 
/PRNewswire/ -- 888poker.com, leader in online poker entertainment, has released an updated mobile application that allows players to now play multi-table tournaments in addition to sit n'gos and cash games.The updated 888poker app is available on all Android powered devices globally.For i Phone users, the following. Tanks - Tank battles in real-time! Application for tanks players online, you can: - Playing with mobile phone and computer - Do a search for your Hall of Fame - Obtain information from the hangar - View the current state of game resources - Share links with friends via E-mail, SMS, social networking Classmates, Vkontakte - Be aware of the latest news of the project Tank is a classic shooter. Play and prohoditi mission. Each mission has a different schemes battle strategies with different enemy tanks. User friendly control panel allows you to control the fire of tanks and construction base. Many types of tanks and armored vehicles, they will turn your phone into Stalingrad. Develop and improve the troops, send tanks into battle and defeat the enemy through superior tactics and firepower! Destroy the enemy! The game is something of Tanchiki era Dendy, the dynamics of modern 3D-action movie, as well as the excitement and thrill of online games. Grandiose tank battles separates you a simple registration. Access to the game is completely free. In the relentless struggle faced live players. Join the battle and lead your troops to glory! Explosive Tank Action times World War! Keywords: wartank tank tanks tanki Tanchiki war tanks wot online on-line on-line world war battle battle strategy.
0 Comments
Underlying all of this is the ISO 20022 metamodel - a model of the models. The UML profile is the metamodel transformed into UML. The metadata is transformed into the syntax of messages used in financial networks. The first syntax supported for messages was XML Schema. ISO 20022 is widely used in financial services. ISO 9362 defines a standard format of Business Identifier Codes (also known as SWIFT-BIC, BIC, SWIFT ID or SWIFT code) approved by the (ISO). It is a unique identification code for both financial and non-financial institutions. The acronym SWIFT stands for the. The ISO has designated SWIFT as the BIC registration authority. When assigned to a non-financial institution, the code may also be known as a Business Entity Identifier or BEI. These codes are used when transferring money between banks, particularly for international, and also for the exchange of other messages between banks. The codes can sometimes be found on account statements. 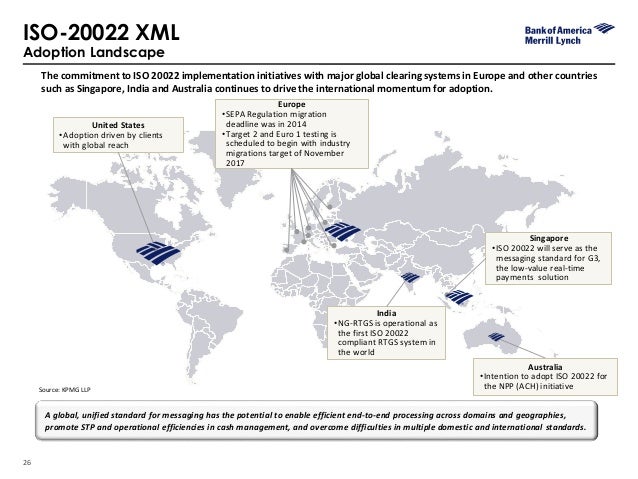
The overlapping issue between ISO 9362 and is discussed in the article (also called IBAN). The SWIFT network does not require a specific format for the transaction so the identification of accounts and transaction types is left to agreements of the transaction partners. In the process of the the European central banks have agreed on a common format based on IBAN and BIC including an XML-based transmission format for standardized transactions; the is a joint gross clearing system in the European Union that does not require the SWIFT network for transmission (see ). The TARGET-directory lists all the BICs of the banks that are attached to the TARGET2-network being a subset of the SWIFT-directory of BICs. Contents • • • • • • • Structure [ ] The previous edition is ISO 9362:2009 (dated 2009-10-01). The SWIFT code is 8 or 11 characters, made up of: • 4 letters: Institution Code or bank code. • 2 letters: country code • 2 letters or digits: location code • if the second character is '0', then it is typically a test BIC as opposed to a BIC used on the live network. • if the second character is '1', then it denotes a passive participant in the SWIFT network • if the second character is '2', then it typically indicates a reverse billing BIC, where the recipient pays for the message as opposed to the more usual mode whereby the sender pays for the message. • 3 letters or digits: branch code, optional ('XXX' for primary office) Where an 8-digit code is given, it may be assumed that it refers to the primary office. SWIFT Standards, a division of The (SWIFT), handles the registration of these codes. Because SWIFT originally introduced what was later standardized as Business Identifier Codes (BICs), they are still often called SWIFT addresses or codes. The 2009 update of ISO 9362 broadened the scope to include non-financial institutions; before then BIC was commonly understood to be an acronym for Bank Identifier Code. There are over 7,500 'live' codes (for partners actively connected to the BIC network) and an estimated 10,000 additional BIC codes which can be used for. 2009 version is now replaced by the latest edition (ISO 9362:2014 dated 2014-12-01). Examples [ ] is an international bank, with its head office in,. The SWIFT code for its primary office is DEUTDEFF: • DEUT identifies Deutsche Bank • DE is the country code for Germany • FF is the code for Frankfurt Deutsche Bank uses an extended code of 11 characters and has assigned branches or processing areas individual extended codes. This allows the payment to be directed to a specific office. For example, DEUTDEFF500 would direct the payment to an office of Deutsche Bank in Bad Homburg. Is a primarily bank, with its head office in. The SWIFT code for its primary office is NEDSZAJJ: • NEDS identifies Nedbank • ZA is the country code for South Africa • JJ is the code for Johannesburg Nedbank has not implemented the extended code of 11 characters and all SWIFT transfers to its accounts are directed to the primary office for processing. Those transfer interfaces that require an 11 digit code would enter NEDSZAJJXXX. Danske Bank is a primarily bank, with its head office in. The SWIFT code for its primary office is DABADKKK: • DABA identifies Danske Bank • DK is the country code for Denmark • KK is the code for Copenhagen. UniCredit Banca is a primarily bank with its head office in. The SWIFT code for its primary office is UNCRITMM: • UNCR identifies Unicredit Banca • IT is the country code for Italy • MM is the code for Milan. Dah Sing Bank is a bank based in Hong Kong that has five branches in mainland China (primary mainland China branch in Shenzhen). The SWIFT code for the branch in Shanghai is DSBACNBXSHA. • DSBA identifies Dah Sing Bank • CN is the country code for China • BXSHA is the code for Shanghai. It uses the 11-digit extended code, and SHA identifies the Shanghai branch. BDO Unibank is the biggest bank in the Philippines, with its head office in Makati. The SWIFT Code for BDO is BNORPHMM. All BDO branches have the same SWIFT Code. • BNOR identifies BDO Unibank • PH is the country code for the Philippines • MM is the code for Metro Manila of which Makati is a part. Note that one bank can seem to have more than one bank identifier in a given country for separation purposes. Bank of East Asia separates its representative branch in the US and its US-based operations for local customers into BEASUS33xxx (following the code used in its home country) and BEAKUS33xxx respectively. This differs from its local mainland China operations which are also BEASCNxxxxx following Hong Kong rather than having a separate identifier code. • An example of this is in the United States. For denominated wires, its SWIFT code is BOFAUS3N. The SWIFT code for wires sent in foreign currency (non-U.S. Dollars) to Bank of America in the United States is BOFAUS6S. In the past, required both BIC and IBAN. Since 2016-02-01 only the IBAN is needed inside the SEPA (European Union and some more countries). In Swift codes consist of 8-11 digits. This example is a swift code for: BSAMLKLXXXX • 'BSAM'- Is the bank code for Sampath Bank • 'LK' Is Sri Lanka's country code • 'LX' is the banks location code. • 'XXX' is the bank's branch code. Because the bank clears its transactions from a central location in Colombo, the letters XXX mean centralised clearing. 12-character SWIFTNet FIN address based on BIC [ ] To identify endpoints on its network, SWIFT also uses 12-character codes that are derived from the BIC of the institution. Such a code consists of the BIC8, followed by 1-character code that identifies the Logical Terminal (LTC), or 'local destination', and the 3-character branch code. These 'BIC12's are not part of the ISO standard, and are only relevant in the context of the messaging platform. Usage [ ] Business Identifier Codes are primarily used for identifying financial and non-financial institutions involving day-to-day business transactions among one or more institutions in transaction life cycle. Example: In SWIFT messages these BICs are embedded within the messages. Let's consider message type for cash transfer MT103, here we can find BIC under different tags like 50a(Ordering Customer), 56a(intermediary), 57a(account with institution), etc. See also [ ] • • (BIN) • (IBAN) • (SWIFT) • () (CIPS) • • • References [ ]. Applies to SAP ERP 5.0 / 6.0 with the focus of One XML format for global payments including SEPA Summary The purpose of this document is: • To provide possible country specific requirements (Europe and Americas) used at Banks to extend the XML format • Development highlights to extend XML format as Global format It is necessary the consultant is aware of all the basic settings of the Automatic payment process. This document is only focused on the creation of complete structure of payment file in XML format with the use of SAP standard template SEPA_CT. Table of Contents 1 Introduction. 2 Benefits of using XML format as Global format 3 General Configuration. 4 XML SEPA Country specific Configuration. 4.1 Rules to determine a SEPA Payment 4.2 General Considerations: All EU – SEPA countries. 4.2.1 Invoice reference in Payment file. 4.3 Country specific changes. 4.3.2 Belgium. 4.3.3 France. 5 XML SEPA extension as Global format 5.1 General Considerations. 5.2 If Beneficiary Bank has ‘IBAN’ 5.3 If Beneficiary has ‘No IBAN’ – User exit 5.4 DKK Payments in Denmark. 5.5 SEK Payments in Sweden. 5.6 NOK Payments in Norway. 5.7 GBP Payments in UK. 5.8 USD Payments in US. 6 Related Content 7 Disclaimer and Liability Notice. 1 Introduction This document aims to understand and to use the XML format as Global format including SEPA requirements / Transactions. To start, SAP’s standard template SEPA_CT for Credit Transfer is used. The basic requirements and technicalities to generate the payment file takes into consideration the requirement of EPC (European Payments Council) The EPC rulebooks contain the business requirements and rules for the operation of the SEPA. The implementation guidelines specify the SEPA core requirements that apply to the UNIFI (ISO20022) XML standards and cover mandatory bank-to-bank messages and optional recommended customer-to-bank messages. 2 Benefits of using XML format as Global format • Converging to SEPA requirements, hence in line with the legal requirements • Less maintenance cost for the company • Adaptability and flexible to accommodate any future changes either by Financial institutions or by business • Highly Secured 3 General Configuration Standard SEPA_CT for credit transfer cannot be used unless changes are adopted as per the bank and country specific requirements. It also depends on the Beneficiary Bank details, Currency of payment and Beneficiary bank country. A new format can be created as there are no formats available for a specific bank/ country. This customization can be either done through modifying the standard function modules of an existing PMW (Payment Medium Workbench) format or through developing via Data Medium Exchange Engine (DMEE). (However this document does not deal with customizing through DMEE). 4 XML SEPA Country specific Configuration ¨ 4.1 Rules to determine a SEPA Payment Data Format • Ordering and beneficiary accounts have to be in IBAN format • BIC is required for both ordering and beneficiary parties How recognize a payment elligible for SEPA The following are few of the rules to make SEPA Credit Transfer: • The transactions have to be in Euro • The Debtor and Creditor must each hold an account with a Participant located within the SEPA • The transfers have to be in the SEPA area. The SEPA area comprises the following countries: o EU: Euro-zone countries: Austria, Belgium, Cyprus, Estonia, France, Finland, Germany, Greece, Ireland, Italy, Luxemburg, Malta, Netherlands, Portugal, Slovakia, Slovenia and Spain o Non-EU: non-Euro-Zone countries Bulgaria, Croatia, Czech Republic, Denmark, Great Britain, Hungary, Latvia, Lithuania, Monaco, Poland, Romania and Sweden 4.2 General Considerations: All EU – SEPA countries SAP Standard template SEPA_CT to be copied as a base template with the necessary naming convention. Path: SPRO IMG Financial Accounting (New) Accounts Receivable and Accounts Payable Business Transactions Outgoing Payments Automatic Outgoing Payments Payment Media Make Settings for Payment Medium Formats from Payment Medium Workbench The above path can also be accessed through the T. Codes OBPM1 / OBPM2 / OBPM3 / OBPM4. The steps in implementing PMW: • Create/ change payment medium formats • Assign variant to the format • Assign the format to payment method Tr. Code: DMEE: To Copy (highlighted below) the Standard template – Fig 1. This will copy all the standard segments of XML format used in payment file. The output structure of standard DMEE file is as shown below – Fig 2 Important Considerations • In the Payment file, a swift BIC code can be on 8 or 11 characters. I.e., for example you can indicate in files as BNPAGB22 or BNPAGB22XXX. The four first characters have to be Letters & the 5 – 6 characters is the ISO country code. • In the payment file, Charges always set-up to SLEV for SEPA payments. This indicates to Bank that default charges set-up will be applied and is born by the debtor. Other charges available for NON SEPA payments are: CRED, DEBT and SHAR. • For example, in Denmark, default charges are shared. You can force charges to what you want by specifying one of the following values: SHAR or DEBT or CRED. This depends on the Business agreement with Beneficiaries. • If the Payment is SEPA relevant, it is always mandatory to have the Payment type information as follows: SEPA • Batch booking option. § If batch booking= true. The bank statement will mention a global debit of the instruction. § If batch booking= false. The bank statement will mention the details of all transactions within the instruction. Sample as follows: 4.2.1 Invoice reference in Payment file Based on the Business requirements, either the payment can be executed by grouping the invoices per vendor & currency or can be separated per invoice. Thus with the following changes in SAP, invoice references can be transferred in the Payment file 4.2.1.1 Group of invoices paid per Vendor & Currency If the invoice is grouped per Vendor & Currency with multiple invoices to pay, it is required to capture all the invoice references in the payment file. This is required as this will serve as the reference to the Beneficiary when the Account statement is received. Code: OBPM2: As per standard format, Note to Payee lines in XML tag can take only 4 reference Atoms that are available (32 * 4 = 128 characters separated by “/”) under XML node. Hence for each vendor, four invoice numbers can only be accommodated in the unstructured field which may not be sufficient for vendors in getting remittance information details in their Account statement. It takes the details based on the above Note to payee text format (Typ 1) line: &FPAYP-XBLNR& Tr.Code: OBPM1 This change is called in the following node in the DMEE file: 4.2.1.2 Reference to be transferred without grouping If grouping is not done, data is transferred with individual invoice references. Hence note to payee length is not a challenge as it is enough to accommodate 18 character length as the invoice reference. 4.3 Country specific changes 4.3.1 Spain Non-resident vendors can be determined based on first three characters from their VAT Reg. “ESN” or Tax number contains pattern “N*” in vendor master control segment. Conditions can be maintained in Regulatory reporting XML tag where amount is greater than 50,000 Euros and Vendor Tax or VAT registration number contains pattern N* or ESN*. Hence the following condition. 4.3.2 Belgium Initiating party ID needs to be mentioned as constant “KBO-BCE” for Belgium 4.3.3 France For French beneficiaries, there is no sort code (it does not exist & sort code is just for UK domestic payments and on 6 digits – Explained below). The details as below: Sample as follows: CCFRFRPPXXX Regulatory Reporting code needs to be mentioned as constant “152” for France where Payer is in France and Payee is outside France Structure in the DMEE output file will appear as Note: Null tags and empty tags are not allowed even on the address field => This involves the rejection of the whole file. Such Tags have to be removed (or fill in) from the file. XML SEPA extension as Global format 5.1 General Considerations XML pain.001.001.02 for SEPA EURO payments and xml iso pain.001.001.03 for Non EURO payments. Thus to facilitate the XML format as Global, it is required to have the header as follows: Sample output at Header f To sum-up: – with pain.001.001.03, it is possible to send SEPA and non SEPA payments – with pain.001.001.02, you can only send SEPA payments 5.2 If Beneficiary Bank has ‘IBAN’ If IBAN is found in the Beneficiary bank account, the IBAN is printed via the structure in the Payment file. If not, BBAN (Basic Bank Account Number) is generated via the user exit Output is: 5.3 If Beneficiary has ‘No IBAN’ – User exit Conditions are put in place to check IBAN, and if blank under segment: Othr, User Exit is triggered. It is required to concatenate Bank Key and account number of the beneficiary under segment Id. Beneficiary Country • Bene. Bank country Currency Type of Transfer US US USD ACH >> CCD or PPD SG US USD ACH >> CCD or PPD SG SG USD NON ACH, can be USD Wires (499) SG CN USD NON ACH, can be USD Wires (499) ** CCD: Corporate Payments, PPD: Personal payments Type of transfer should appear under tag Cd as follows: • For any USD local payments, service level code is NURG as follows NURG • For any USD foreign payments, service level code is URGP as follows Sample as follows: TRF false 3 950.00 URGP 2014-07-06 Also, the formatting for the debtor account tag should be Sample as follows: – – 30XXXX84 – BBAN USD 6. Related Content For more information for DMEE payment structure and configuration details, visit SAP help: 7. Disclaimer and Liability Notice This document may discuss sample coding or other information that does not include SAP official interfaces and therefore is not supported by SAP. Changes made based on this information are not supported and can be overwritten during an upgrade. 
Autodesk 3ds Max 2015, 128G1. Autodesk 3ds Max Design 2015, 495G1. Autodesk 3ds Max Entertainment Creation Suite Premium 2015, 774G1. Autodesk 3ds Max Entertainment Creation Suite Standard 2015, 661G1. Autodesk 3ds Max with SoftImage 2015, 978G1. Autodesk Advance Concrete 2015, 960G1. This Service Pack contains recent fixes for Autodesk® 3ds Max 2015 software. It is strongly recommended that you read the readme document before installing the software. For future reference, you should save the readme to your hard drive. Download 3ds Max 2015 SP3 (exe - 189 Mb) Readme • • • • • • If the self-extracting EXE file above fails to properly install, please try to install directly with the Microsoft Installer Patch (MSP) file below. If you continue to encounter problems, please contact Product Support. (msp - 189 Mb). 4K Video Downloader 3.8 Crack is very powerful software to download video, audio and subtitles from YouTube in high quality and as fast. If any one went to video on your iPad, iPhone. I use this software in my windows operating system 32 bit and 64 bit. I think you search many place but many site not give you best download link. Here our team give you best download link then you can easily use this software in your PC. So download our link and enjoy this software. 4K Video Downloader Features: It software download entire playlists and channels from. Abstract The distinct lifestyle of obligately intracellular bacteria can alter fundamental forces that drive and constrain genome change. In this study, sequencing the 792-kb genome of Blochmannia pennsylvanicus, an obligate endosymbiont of Camponotus pennsylvanicus, enabled us to trace evolutionary changes that occurred in the context of a bacterial–ant association. Comparison to the genome of Blochmannia floridanus reveals differential loss of genes involved in cofactor biosynthesis, the composition and structure of the cell wall and membrane, gene regulation, and DNA replication. However, the two Blochmannia species show complete conservation in the order and strand orientation of shared genes. 2002) and near-perfect synteny since the establishment of this association 150–200 million years ago (Mya) (van Ham et al. This exceptional stasis of genome architecture contrasts with lability of free-living and parasitic bacterial genomes. Genome stability may. Download as PowerPoint Slide.  
This finding of extreme stasis in genome architecture, also reported previously for the aphid endosymbiont Buchnera, suggests that genome stability characterizes long-term bacterial mutualists of insects and constrains their evolutionary potential. Genome-wide analyses of protein divergences reveal 10- to 50-fold faster amino acid substitution rates in Blochmannia compared to related bacteria. Despite these varying features of genome evolution, a striking correlation in the relative divergences of proteins indicates parallel functional constraints on gene functions across ecologically distinct bacterial groups. Furthermore, the increased rates of amino acid substitution and gene loss in Blochmannia have occurred in a lineage-specific fashion, which may reflect life history differences of their ant hosts. Genome sequencing provides a rich data set to predict the metabolic capabilities of organisms, and comparative analyses among closely related species offer a powerful approach to examine mechanisms of genome flux. Since 2000, genomics has shed light on the metabolism and evolution of obligately intracellular, mutualistic bacteria that have coevolved with various insect groups for tens to hundreds of millions of years (). Fully sequenced genomes of Buchnera associated with aphids (;; ), Wigglesworthia of tsetse flies (), and Blochmannia associated with Camponotus () are extremely streamlined yet retain basic cellular processes and specific biosynthetic abilities required by the insect host. Genome comparisons within Buchnera have shown stability, with no gene acquisition, inversions, or translocations throughout 50–70 million years (Myr) of evolution within aphids () and near-perfect synteny since the establishment of this association 150–200 million years ago (Mya) (). This exceptional stasis of genome architecture contrasts with lability of free-living and parasitic bacterial genomes. Genome stability may reflect the dearth of molecular tools for gene exchange (e.g., phage, certain rec genes, and repeated DNA sequences) in this mutualist and limited ecological opportunities to recombine with genetically distinct bacteria (;; ). Such constraints on genome changes in stable mutualists may profoundly affect the evolutionary potential of these bacteria and their hosts. However, owing to the lack of multiple sequenced genomes within endosymbiont groups, genome stability in other long-term endosymbionts has remained untested. In order to contribute to a more comprehensive model of genome evolution in ancient endosymbiotic associations, we have evaluated genome dynamics in Blochmannia, a bacterial genus that is closely related to Buchnera and has cospeciated with ants for ∼30 Myr. The wide range of interactions between ants and other species, including plants, fungi, trophobionts, other insects, and diverse bacteria (;; ), may explain the huge ecological success of ants, which play dominant roles in nutrient turnover in terrestrial ecosystems and include more than twice as many species as mammals (). Blochmannia is the most evolutionarily stable ant associate and lives exclusively within cells of the closely related genera Polyrhachis, Colobopsis, and Camponotus (;;;; ). Blochmannia has been studied most extensively in Camponotus, the second largest ant genus, with ∼1000 species () ranging from omnivores to specialists on plant secretions and homopteran exudates (;;; Davidson, ) and with nesting habitats including wood, soil beneath rocks, and the rainforest canopy (). The 706-kb sequence of B. Floridanus ( Blochmannia of host Camponotus floridanus) indicated this ant endosymbiont retains numerous metabolic pathways that may be involved in host nutrition, including nitrogen recycling and assimilation, biosynthesis of amino acids and fatty acids, and sulfate reduction (). Blochmannia is thought to be important during host development (; ), but its specific roles in host physiology and ecology and its functional variation across ant species remain unclear. We have sequenced the 792-kb genome of Blochmannia pennsylvanicus ( Blochmannia associated with Camponotus pennsylvanicus) to examine genome changes since this lineage and B. Floridanus diverged from a common ancestor ∼16–20 Mya (). Comparing genome inventories and architectures of the two Blochmannia strains allowed us to trace functional and structural changes that occurred in the context of this bacterium–ant interaction and to contrast genome dynamics and protein evolution in two mutualist groups ( Blochmannia and Buchnera). Results Genome features The B. Pennsylvanicus genome consists of a 791,654-bp circular chromosome that we sequenced to 12× coverage (;; Supplemental Fig. S1; GenBank entry CP000016). This AT-rich genome has a relatively low percentage of coding DNA (76.7%) and shows a GC skew that distinguishes the leading and the lagging strands of replication [(G–C)/(G+C) averaged across 2.5-kb sliding windows]. We identified predicted ribosomal binding sites (RBSs) for 479 of 610 open reading frames (ORFs), but only 42 had the canonical AGGAG motif. Surprisingly, two pseudogenes ( rpmD and uvrD) had detectable RBSs. Differential gene loss, yet complete stability of genome architecture within Blochmannia The 86-kb size difference between the Blochmannia genomes ( B. Pennsylvanicus, 792 kb; B. Floridanus, 706 kb) largely reflects differential gene loss between the two lineages, with a greater extent of loss in B. Both genomes possess several intact genes that are missing or pseudogenes in the other genome (). Although lateral gene transfer could, in principle, account for genes distinct to one Blochmannia genome, all genes grouped closely with enterobacterial homologs and showed amino acid and codon usage patterns typical of their resident genome. Notably, the order and strand orientation (i.e., genome architecture) is completely conserved for their 635 shared genes, including cases in which a functional gene in one genome is a pseudogene in the second. Assuming that the common ancestor of the two Blochmannia strains encoded at least their combined set of 615 ORFs, then gene loss or inactivation in the lineage leading to B. Floridanus occurred at an approximate rate of one ORF per 0.64–0.8 Myr [(25 ORFs lost)/(16 Myr) to (25 ORFs lost)/(20 Myr)]. Gene loss in the B. Pennsylvanicus lineage is apparently ∼6.5 times slower, with loss or inactivation of one ORF per 4.0–5.0 Myr [(4 ORFs lost)/(16 Myr) to (4 ORFs lost)/(20 Myr)]. Estimated rates of gene loss in B. Floridanus exceed those in Buchnera, in which one ORF was lost or inactivated per 2.70–3.60 Myr between Buchnera–B. Pistaciae and Buchnera–Acyrthosiphon pisum [(111 ORFs lost)/(150 Myr × 2) to (111 ORFs lost)/(200 Myr × 2)] and per 1.70–2.38 Myr between Buchnera–A. Pisum and Buchnera–Schizaphis graminum [(59 ORFs lost)/(50 Myr × 2) to (59 ORFs lost)/(70 Myr × 2)]. Rates of loss may be underestimated if the same genes were deleted independently along lineages. Mechanisms that influence rates of gene loss may include changes in the strength and/or efficacy of selection to maintain genes, as well as differences in underlying rates of gene knockouts and deletion (see Discussion). Truncations and frameshifts in otherwise conserved Blochmannia genes In all, 13 Blochmannia genes show a significant (20%–40%) truncation compared to Escherichia coli orthologs but apparently encode functional proteins. These genes include aceF, aroK, aroQ, ftsK, ftsY, hfq, mreC, pheA, rpoZ, thrS, trpD, yfcB, and yqeI. In all but one case ( hfq, see below), truncations are shared by B. Pennsylvanicus and B. Floridanus and thus likely occurred before the divergence of these lineages. Certain truncations are also shared between Blochmannia and Wigglesworthia ( yfcB, rpoZ, ftsK), Buchnera ( aroQ), or among all three mutualist groups ( aceF, ftsY). Although a >20% length reduction is often interpreted as evidence for loss of gene function (e.g., ), the sequence conservation of truncated Blochmannia genes suggests they encode functional proteins. First, nonsynonymous divergence ( dN) between B. Pennsylvanicus and B. Floridanus is relatively low at the 12 truncated genes they share (average of 0.3012 ± 0.14), whereas synonymous divergence ( dS) exceeds 2 for each gene. Although dN/ dS is difficult to calculate because of saturation of synonymous sites, this ratio is below 0.13 for each gene. Moreover, apart from RpoZ, protein divergences at truncated genes are generally comparable to other ORFs (average protein divergence of 0.569 ± 0.281, compared to the genome-wide average of 0.565 ± 0.309). RpoZ is quite divergent (2.16), but the fact that dN/ dS ≪ 1 suggests the truncation did not eliminate gene function. The one truncated gene ( hfq) in B. Pennsylvanicus that lacks an ortholog in B. Floridanus has a relatively low protein divergence with the closest outgroup, Wigglesworthia (0.576). Thus, in the absence of detailed biochemical and structural information for each of these genes, a 20% truncation appears too conservative a criterion for inferring loss of function. In Blochmannia, truncated genes clearly differ from annotated pseudogenes, in which frameshift and nonsense mutations introduce multiple stop codons throughout the gene. Circular map of the B. Pennsylvanicus genome and genome features. The origin of replication was putatively set upstream of gidA, in accordance with the shift of GC skew. The concentric rings denote the following features: (1) numbered base pair coordinates beginning with base one of the gidA open reading frame (ORF); (2) GC skew as calculated by (G–C)/(G+C) using a 2.5-kb sliding window; (3) ORFS present on the leading strand (+); (4) ORFS present on the lagging strand (–); (5) pseudogenes of B. Pennsylvanicus that are present in (orange) or absent from (black) B. Floridanus; (6) ORFS of B. Pennsylvanicus that are shared with (red), pseudogenes in (yellow), or absent in (dark blue) the genome of B. Floridanus; (7) transfer RNAs (tRNAs) that are shared with (gray) or absent in (pink) B. Floridanus; (8) ribosomal RNAs (rRNAs) 5S, 16S, and 23S (purple). Circular plot and GC skew analysis were generated by the JENA Prokaryotic Genome Viewer–Export Version v3.06.04. We also detected a single, short frameshift in each of B. Pennsylvanicus ytfM, ybiS, hisH, and ubiF. The first two genes are unclassified proteins, while the latter are required for histidine and ubiquinone biosynthesis, respectively (see legend). Each indel occurred within a 9–11-bp string of consecutive As (). Reanalysis of B. Floridanus pseudogenes revealed a similar pattern at ytfM, ybiS, and gmhB, in which single frameshifts occur within poly(A) or poly(T) tracts. Poly(A) tracts are very common within Blochmannia ORFs (e.g., B. Pennsylvanicus has 133 poly(A) or (T) tracts >9 bp long) and likely reflect AT mutational bias and reduced selective pressures. The indels in ytfM, ybiS, hisH, and ubiF each show high coverage in the B. Pennsylvanicus shotgun assembly (5–15 independent clones). In the case of hisH, the frameshift occurs across independent PCR products sequenced directly with varying types of sequencing chemistry. Despite their consistency, we hesitate to interpret these frameshifts as firm evidence of pseudogenes. Notably, apart from a single frameshift, these genes would otherwise encode intact proteins that are relatively conserved between the two Blochmannia genomes, with an average protein divergence (0.67 ± 0.16) that is only slightly above the average for other ORFs (0.565 ± 0.309). The occurrence of frameshifts within homopolymeric tracks is consistent with slippage during transcription (; ) or during translation (; ) that could restore the full-length protein (see Discussion). If the frameshifts have, indeed, disrupted protein functions, these mutations must have occurred very recently since rapid mutation in Blochmannia () is expected to erode pseudogenes quickly. Metabolic similarities of Blochmannia spp. Analysis of B. Pennsylvanicus and reanalysis of other insect mutualist genomes using MultiFun () allowed more comprehensive metabolic comparisons across groups (see and Supplemental Tables S1 and S2). The two Blochmannia genomes share nearly all metabolic pathways thought to contribute to host nutrition, including most biosynthetic functions, sulfate assimilation and metabolism, and hydrolysis of urea (see ). Metabolic differences between Blochmannia spp. The 30 ORFs that distinguish the two Blochmannia genomes have a range of predicted cellular functions that may alter host–symbiont metabolic exchanges (Figs. Pennsylvanicus retains coaADE and dfp for the biosynthesis of coenzyme A, an essential cofactor and substrate for the TCA cycle (). In contrast, the deletion of these genes from B. Floridanus implies that this endosymbiont does not require coenzyme A, uses other enzymes for its synthesis, or imports this cofactor from the host. Several differences between the Blochmannia genomes involve the biosynthesis, transport, and mediation of cellular wall and membrane components. Pennsylvanicus retains six distinct ORFs that contribute to the de novo synthesis of peptidoglycan (murein), the major constituent of Gram-negative bacterial cell walls (). In addition to MurI, it retains the complete pathway for the biosynthesis of isoprenoids (), essential substrates for synthesis of peptidoglycan (), and several organic compounds including carotinoids, glycosyl carrier lipids, and the side chains of ubiquinone (; ). Floridanus, the absence of murI and the interruption of isoprenoid biosynthesis implies this bacterium may import both D-glutamate and trans, trans-farnesyl diphosphate (isoprenoid precursor) from the host, might synthesize these components using other enzymes, or produce a cell wall lacking these structural elements (i.e., isoprenoids). Comparison of B. Pennsylvanicus and B. Floridanus gene contents. Genes in the outer section of each circle are unique to one genome, while those listed in the intersection of the two circles are shared. Note that seven shared genes are apparently functional in one genome, but pseudogenes in the other. The truncation of B. Floridanus dnaX (see text) is not included as a pseudogene here. The fusion of yidCD in B. Pennsylvanicus is counted as two intact genes. (*) The 628 shared intact genes include genes that have a single frameshift within long poly(A) tracts but are otherwise intact (see text). These loci include B. Pennsylvanicus hisH, ytfM, ubiF, and ybiS and B. Floridanus ytfM, ybiS, and gmhB. Pennsylvanicus is the first fully sequenced insect mutualist that retains the entire sec-dependent secretory pathway, including the chaperonin SecB. This general pathway mediates the export and translocation of numerous proteins to the periplasm or inner membrane (). The other insect mutualist genomes lack certain components (typically secBDF) of this pathway (;;;; ), although these losses are not expected to eliminate function (). Pennsylvanicus also retains the inner-membrane-bound heat-induced protease HtpX and the periplasmic chaperonins LolB and DsbA. Together, these results suggest that B. Pennsylvanicus is better able to respond to cellular stress and ensure proper transport localization, conformation, and decomposition of gene products. Distinct membrane features in B. Pennsylvanicus also include its retention of the inner membrane proteins YciC, putatively involved in transport, and BacA, which confers bacitracin resistance. While insect mutualists have lost many regulatory genes, B. Pennsylvanicus encodes three that are missing from B. Floridanus ( hns, hfq, and mntR). Floridanus and Wigglesworthia, B. Pennsylvanicus lacks DnaA, considered important in the initiation of DNA replication (; ). Floridanus, the HU-like nucleoprotein HlpA might be involved in starting the nucleosome (), and in B. Pennsylvanicus, the nucleoprotein Hns might also be recruited to the replication origin for this purpose. Neither Blochmannia genome encodes the θ subunit of the holoenzyme DNA polymerase III, but B. Pennsylvanicus retains the τ and γ subunits of DnaX. The τ subunit acts as a molecular tether that couples DnaB (DNA helicase) to the core of DNA polymerase III (α and ϵ) as the replication fork progresses from the origin (; ). Thus, the inability of B. Floridanus, Buchnera, and Wigglesworthia to transcribe the τ subunit is expected to decrease the efficiency, accuracy, and processivity of the holoenzyme (). These distinct features of the B. Pennsylvanicus replication machinery might contribute to the apparent 29-kb shift observed in B. Floridanus GC skew relative to B. Pennsylvanicus (see Supplemental Fig. Long intergenic spacers in B. Pennsylvanicus Intergenic spacers in B. Pennsylvanicus are significantly longer (average 291 bp) than homologous spacers in B. Floridanus (average 180 bp; Wilcoxon Rank sum test, p 1 kb) in B. Multiple gene-finding tools did not detect ORFs or recognizable pseudogenes in any spacers, which have lower GC contents (average of 20.0%) compared to ORFs (average of 32.1%) () and may represent eroded pseudogenes and/or regions involved in gene regulation. Polymorphism within B. Pennsylvanicus Polymorphisms in the pooled symbiont population used for library construction were detectable as well-supported (Phred scores >40) discrepancies in the genome assembly that were represented by at least two independent clones. Nearly all (445/497) polymorphisms are single nucleotide polymorphisms (SNPs). Those located within ORFs occur primarily at third codon positions (). Although the majority of SNPs are located within ORFs, a disproportionate number of SNPs (35%) and indels (96% of insertions and/or deletions) occur within the intergenic regions (which comprise 23% of the B. Pennsylvanicus genome). The two polymorphic indels within ORFs produce amino acid insertions/deletions in sucA and aroE. Comparison of protein divergences Wide variation in protein divergence across loci indicates variable functional constraint across Blochmannia proteins (; Supplemental Tables S3 and S5). Amino acid biosynthetic genes, ribosomal proteins, and particular chaperonins are exceptionally conserved (a mean protein divergence of 0.448, with a 99% confidence interval of 0.400–0.504), while surface structures [mean of 0.725 (0.533–0.991)] and hypothetical or unclassified proteins [mean of 0.775 (0.667–0.898)] are quite divergent (Supplemental Table S5). The membrane proteins tolA and tonB are particularly divergent (divergences of 1.74 and 2.64, respectively) (Supplemental Table S3). Previous studies have demonstrated a negative relationship between GC content of endosymbiont genes and their level of divergence from free-living relatives (; ), suggesting that amino acid changes in proteins under strong functional constraint are less severely affected by AT mutational bias. A strong negative association between GC content and protein divergence in Blochmannia (Supplemental Fig. S3) indicates this relationship also holds when protein divergences are estimated within a mutualist group (rather than to a more distant free-living relative). Protein divergences were, on average, ∼1.88 times faster in B. Floridanus lineage compared to B. Pennsylvanicus lineage. Genes with particularly elevated rates in B. Floridanus include secG, rpsJ, rpsR, and rnpA, each of which evolves more than 10-fold faster in B. Floridanus compared to B. Pennsylvanicus (; Supplemental Table S4). A 14-fold rate acceleration at ispE may reflect the loss or disruption of other genes involved in isoprenoid biosynthesis in the B. Floridanus genome and relaxed selection at this remaining isp gene. Sequence context of homopolymeric frameshift mutations within five Blochmannia genes. The single frameshifts in Blochmannia genes ( A) hisH, ( B) ubiF, ( C) ybiS, ( D) ytfM, and ( E) gmhB were “corrected” manually, based on comparisons with sequences that lack the frameshift. Genes were then aligned by their inferred amino acid sequence against E. Underlined nucleotides indicate poly(A) and poly(T) tracts in which frameshifts occur. Regions highlighted in gray indicate the putative sites of transcriptional or translational slippage that may restore the proper reading frame. (•) Amino acid identical to the top sequence ( B. Pennsylvanicus genome offered the first opportunity to compare genome-wide patterns of protein evolution in the context of distinct endosymbiotic associations. A previous study showed accelerated evolution at 16S rDNA and at synonymous positions of select Blochmannia genes compared to enteric bacteria and even compared to the rapidly evolving Buchnera (). Here, we tested whether the Blochmannia genome also undergoes exceptionally fast rates of protein evolution. Strong correlations in divergences at homologous genes indicate parallel functional constraints in Blochmannia compared to Buchnera and E. Coli– Photorhabdus luminescens (). However, notably higher protein divergences between Blochmannia spp., despite their relatively recent split, reflects a several-fold acceleration in absolute rates of protein evolution. Among the 302 genes for which RSD detected orthologs across Blochmannia and Buchnera and estimated divergences within each mutualist pair, the average substitution rate in Blochmannia was 0.0132 to 0.0166 amino acid substitutions/site per million years (based on a 20- to 16-Myr divergence, respectively). This is ∼10 times faster than rates in Buchnera for the same gene set (0.00258–0.00362 amino acid substitutions/site per million years, based on a 70–50-Myr divergence). Previous studies have shown that nonsynonymous sites in Buchnera evolve an average of twofold () and up to 10-fold () faster than in the enterics. Likewise, we found an average ∼50-fold rate acceleration in Blochmannia compared to E. Coli– Salmonella typhimurium across the 538 genes for which RSD detected orthologs (0.0142–0.0177 in Blochmannia, and 0.000245–0.000367 for E. Typhimurium assuming a divergence time of 150–100 Myr). Deviations from this general trend highlight genes that may experience different levels of functional constraint among genomes. For example, relative to other genes in the genome, amino acid biosynthetic genes evolve slightly slower in Blochmannia than in the enterics, while genes involved in translation (comprised largely of ribosomal proteins) evolve slightly faster than expected (). Comparisons of average protein divergences within and between major functional categories confirmed many of these observations above (Supplemental Table S5). Blochmannia, Buchnera, and E. Luminescens show similar relative divergences across most functional categories, such as relatively high divergences of unclassified and hypothetical genes, loci for surface structures, cell membrane components; moderate divergences of genes encoding cofactor biosynthesis, information transfer, cell processes, metabolism, information transfer; and relatively low divergences of genes for nucleotide biosynthesis and amino acid biosynthesis. Striking differences among pairs include relatively high divergence of chaperonins and fatty acid biosynthetic genes in Buchnera, the relative conservation of regulation genes in both endosymbionts, and the slightly higher divergence of translation genes in Blochmannia, as noted above. Discussion In contrast to many pathogenic and free-living bacterial species in which lateral gene transfer, chromosomal inversions, and translocations drive changes in gene order and content, we found complete conservation in gene order and orientation between two genomes of Blochmannia that diverged 16–20 Mya. This exceptional genome stability, first demonstrated in Buchnera (; ), suggests that Blochmannia also lacks genetic machinery for gene inversions or translocation. Notably, like Buchnera, both Blochmannia strains lack RecA, numerous other recombination functions (RadA and Rec-FOR), phage, and have relatively low levels of repeated DNA. The gene content of B. Pennsylvanicus differs from B. Floridanus only by 3.6% (24/659); however, genes specifically retained in B. Pennsylvanicus span varied functional categories that may affect its metabolic capabilities and host interaction. The retention of genes for cell wall integrity, chaperonins, gene regulation, and DNA replication in B. Pennsylvanicus may reflect different bacterial requirements for the maintenance of cell processes and structures within C. Pennsylvanicus bacteriocytes. Furthermore, the ability to synthesize both coenzyme A and isoprenoids likely benefits B. Pennsylvanicus and their ant hosts. Genome stasis implies gene losses in either Blochmannia lineage are irreversible, such that deletions of metabolic functions may constrain the evolutionary potential of this association. Such constraints have been proposed in Buchnera (), where the loss of genes for sulfur reduction and cysteine biosynthesis in Buchnera–SG may constrain the S. Graminum host to its relatively cysteine-rich grass diet. Certain aspects of Blochmannia metabolism remain unclear, owing to the uncertainty of whether or not single frameshifts in particular genes eliminate function. For example, if single indels within poly(A) tracts of hisH and ubiF are subject to correction of some type, the encoded proteins might be functional in B. One possible mechanism for correction may be instability of such frameshifts during DNA replication, such that populations include heterogeneous genomes with different numbers of adenines in the homopolymeric repeats (e.g., ). However, among the many C. Pennsylvanicus colonies used for symbiont library construction, we found no evidence for variation in lengths of these or any homopolymeric tracts. Alternatively, transcriptional slippage, or “stuttering” within repeat mononucleotides is well-documented in E. Coli (), where it often occurs within poly(A) or poly(T) tracts (). In a survey of published bacterial genomes, Baranov et al. () identified several “pseudo pseudogenes,” for which transcriptional slippage could correct a frameshift within poly(A) or poly(T) tracts and restore uninterrupted ORFs. These authors describe a mechanism in which the RNA chain dissociates from the DNA template and reassociates in a new location. Third, functional proteins may be restored by frame-shifting during translation, or “recoding,” a phenomenon that occurs in yeast (), archea (), and E. Coli () and can also follow poly(A) tracts, especially in the form of A_AAA_AAG (Baranov et al., ). Such slippage has been proposed to explain aberrant indels in animal mitochondrial genes (), and in principle, might operate in endosymbionts. Given the multiple levels at which frameshifts within homopolymeric sequences may be corrected, we argue that these mutations should be interpreted cautiously, and with consideration of whether the gene otherwise encodes a full-length ORF. Although it is possible that such genes represent very recent loss of function, we take the approach of Baranov et al. () in questioning whether such genes should be annotated as pseudogenes. Conservation of genome architecture and overall similarity in gene content within Blochmannia contrasts with the exceptionally fast rates of sequence evolution observed in this group. Namely, protein divergences are higher for the two ant mutualists than within much older bacterial pairs. A previous study showed that amino acid changes were influenced by AT mutational pressure to a greater extent in basal lineages in Buchnera, suggesting endosymbiont proteins were more tolerant of these presumably deleterious changes early in the association (). By the same token, rapid rates of protein evolution in Blochmannia might reflect the younger age of this association (∼30 Myr, compared to ∼150–200 Myr for Buchnera). Despite an overall rate acceleration in Blochmannia, the relative levels of divergence among proteins were strikingly similar to those of Buchnera and the enterobacteria. Although different types of selection shape free-living and endosymbiotic bacteria, this observed correlation suggests parallel functional constraints across many shared proteins. Comparison of intergenic spacers (IGSs) in B. Pennsylvanicus and B. ( A) A significant relationship in the lengths of homologous spacers implies a certain conservation of spacer length (Spearman's ρ = 0.7895; p. A deviation from this pattern occurs in Blochmannia, where genes involved in translation are not the most conserved functional category relative to the enterics (Supplemental Table S5). We compared the ribosomal proteins that deviated from the best fit line in and those that showed a significant acceleration in the B. Floridanus lineage (; Supplemental Table S4) to the detailed crystal structures of the 30S and 50S ribosomal subunits (; ). These subunits assemble from 52 ribosomal proteins that form scaffolds around the 5S, 16S, and 23S ribosomal RNAs. Notably, many of the genes that show elevated substitution rates are positioned at the periphery of the ribosomal subunits, well away from the active regions at their interface. Furthermore, those proteins specifically involved the initiation and aggregation of the 30S and 50S ribosomes (RplL, RpsG) and those that have regulatory functions (RplADKT, RpsH), while exhibiting elevated rates were not among the genes with the highest divergences. These fast rates of substitution at ribosomal proteins might be influenced by several factors, including the effects of genome-wide patterns of nucleotide substitution; changes in regulatory interactions; compensatory changes due to the AT bias and consequently the secondary structure of the 16S, 23S, and 5S rRNAs; and the inactivation of the rpmD large ribosomal subunit locus. These amino acid substitutions in the ribosomal subunits might reduce the stability of the ribosome and thus decrease translation efficiency. Within Blochmannia, faster divergence in the B. Floridanus lineage at nearly all (∼90%) proteins may reflect elevated mutation rates, reduced selective coefficients, or smaller effective population size of this symbiont and/or its host with associated increased genetic drift. Although data to distinguish these alternatives are limited, an analysis of four gene regions ( groEL, rpsB, atpB, and gidA, all of which evolve faster in the B. Floridanus than B. Pennsylvanicus lineage) showed no consistent increase in dN/ dS in B. Floridanus or its close relatives () that would be predicted under relaxed selection or drift hypotheses. Evidence supporting the mutation hypothesis includes a lower genomic GC content for B. Floridanus than B. Pennsylvanicus (). Because the two Blochmannia genomes have the same set of DNA repair genes, there is no a priori reason to propose that B. Floridanus has a faster rate of mutations per replication. However, the year-round activity of C. Floridanus and its relatives in the subgenus Myrmothrix contrasts with the winter dormancy of C. Pennsylvanicus and related temperate species in the subgenus Camponotus. This activity may increase the number of host and bacterial generations per year and, consequently, the rate of mutations per unit time. Likewise, a combination of elevated mutation, relaxed selection, and/or increased genetic drift in the B. Floridanus lineage may account for faster rates of gene loss compared to the B. Pennsylvanicus lineage (see Lawrence and Roth [] for theoretical framework). However, given that current data cannot distinguish points along the B. Floridanus lineage at which evolutionary rates accelerated or at which particular genes were lost, at this time we cannot link these changes to specific aspects of host ecology. More intensive sampling of Blochmannia across ecologically diverse hosts should allow such connections to be made. Correlated protein divergences suggest parallel selective constraints in endosymbionts and free-living bacteria. Protein divergence in Blochmannia shows a strong correlation with divergence at orthologous genes of ( A) Buchnera (– A. Pisum versus – S. Graminum) and ( B) E. Coli versus P. Lines that best fit the data and intercept zero are shown, but significance was tested using the nonparametric test of association (JMP 4.0; SAS Institute Inc.). For all bacterial pairs, the most conserved genes include certain chaperonins and translation functions (mostly ribosomal proteins), while the most divergent genes include surface structures. Substitution rates above 2 are prone to saturation but are included here for comparison. Supplemental Table S3 lists all of the pairwise divergences. Given the wide variation in Camponotus nutritional ecology, ranging from plant-specialists to omnivorous species, it seems unlikely that a single nutrient is lacking in the diet of all species that house Blochmannia. Rather than supplementing specific dietary deficiencies, nutritional functions of Blochmannia may play critical roles during two “starvation” phases of the host when metabolic demands exceed the available food supply—metamorphosis and colony founding (). Recent work has shown that Blochmannia proliferate during pupation (), a stage of metamorphosis when the host must construct all components of the adult body plan with no food intake (). Genome sequence data provide a starting point for experimental analyses to clarify the functional significance of this mutualism, to explore the implications of genome variability on the physiology and ecology of both symbiotic partners, and to clarify the levels and timing of selection that shape this long-term bacterium–ant association. Methods Blochmannia genome sequencing and assembly B. Pennsylvanicus genomic DNA (gDNA) was prepared from worker and larvae C. Pennsylvanicus collected from five colonies at two sites in Falmouth, Massachusetts, USA. The gDNA was either extracted directly from the agarose plugs containing the purified bacterial cells () or gelpurified from a chromosomal fragment resolved through Pulsed Field Gel Electrophoresis (PFGE) (). Short (1.5–2.5 kb) insert libraries were generated from hydrosheared DNA using a double adaptor kit (SeqWright Inc.) (). Plasmid clones were purified and bidirectionally sequenced using BigDye v3.0 chemistry on either an ABI3700 or an ABI3730xl (Applied Biosystems). Detailed methods for library construction and sequencing are provided in the Supplemental material. Accelerated rates of evolution in the lineage leading to B. Floridanus compared to B. Pennsylvanicus, since their divergence from a common ancestor. Rates of protein divergence were compared using a relative rates test with E. Coli as an outgroup (see Methods). The analysis included 516 proteins for which RSD identified orthologs in the two genomes, and for which divergences between the two Blochmannia or between either endosymbiont and E. Coli did not exceed 2.0. On average, proteins evolved 1.88 times faster in B. Floridanus compared to in B. Only 50 of the proteins tested evolved more slowly in B. Pennsylvanicus than in B. Floridanus or at the same rate, with values ≤1on the histogram, while 467 genes evolved faster in B. Supplemental Table S4 lists proteins with particularly accelerated evolutionary rates in B. Raw sequence data were analyzed by PHRED (; ) () and screened using BLASTN/X () for ant host contamination. Sequence reads that were putatively identified as γ-Proteobacterial ( E ≤ 1 –10) were assembled using ARACHNE 2 () (). The resulting contigs were analyzed by hand in CONSED () and using BAMBUS (). Pennsylvanicus assembly was aligned using LAGAN () () to the published B. Floridanus genome (NC005061), which facilitated primer design for gap closure by PCR. Annotation and metabolic reconstruction Open reading frames (ORFs) were identified iteratively using GLIMMER v2.10a () () and gene orthology predictions based on BLASTP sequence similarity to the NR, SWISS-PROT, and ECOLI databases; HMMR searches against Pfam_ls (); and identification of E. Coli orthologs using the Reciprocal Sequence Distance (RSD) program () (details of the RSD method are noted below under “Comparison of protein divergences”). The few discrepancies among methods were limited to cases of differential loss of one gene from a pair of paralogs ( ilvBG, tufAB, argFI), the presence of gene fusions ( yidCD) or split genes ( trpDG), or failure of RSD to identify a given ortholog because of high sequence divergence. Three pseudogenes ( uvrD, yqiC, rpmD) were detected as regions with similarity to functional ORFs in other genomes, but with multiple indels and missense mutations resulting in stop codons throughout each gene. The two most degraded pseudogenes ( rpmD and yqiC) were undetectable by BLASTX and were only identified because of the conservation of gene order between the two Blochmannia genomes. Among the three pseudogenes, uvrD retains the longest (107 amino acids) intact reading frame with similarity to functional orthologs, but even this region is just 15% of the length of UvrD in E. Coli and other outgroups. In this sense, annotated pseudogenes clearly differ from truncated B. Pennsylvanicus ORFs, which retain at least 60% of the length of orthologous proteins. Pennsylvanicus pseudogenes also differ from genes with single frameshifts within homopolymeric regions ( hisH, ubiF, ytfM, ybiS), since the latter would encode intact, relatively conserved proteins if the frame-shift were corrected by slippage during transcription or translation (see, Results, and Discussion). ORFS and RNAs were manually curated using a Generic Model Organism Database (GMOD) Web browser. ORFs that lacked sequence similarity to any entry in GenBank or the Comprehensive Microbial Resource () and lacked any predicted protein domains in Pfam_ls were excluded from the annotation. Functional and pseudo-transfer RNAs (tRNAs) were identified using tRNAscan-SE (), and ribosomal RNAs and structural RNAs were identified by BLASTN searches of the intergenic regions. Blochmannia gene functions and interactions were inferred from orthologs of E. Coli K12 MG1655 described in GenProtEC () () and characterized by MultiFun (), two resources that represent functions of ∼80% of the 4401 genes in E. Metabolic pathways were evaluated using the reference pathways available for E. Coli at EcoCyc () and KEGG (). Genomes of other insect mutualists were reanalyzed in the same manner for a consistent metabolic comparison. Comparison of protein divergences The RSD algorithm () was used to identify the reciprocal best BLAST hits (rbh) between translated ORFs of select bacterial genomes. The program used BLAST to identify potential matches of a given translated gene, aligned all potential matches using CLUSTALW (), and calculated a maximum likelihood estimation of amino acid substitutions between proteins using PAML (). Protein divergences were based on an empirical amino acid substitution rate matrix () and accounted for variation in evolutionary rates among protein sites using a γ distribution with shape parameter α = 1.53 (as recommended by ). The protein with the lowest divergence was then BLASTed against the first genome, followed by the alignment and divergence calculations. If the protein match with the lowest divergence was the same as the original query sequence, the pair was considered orthologous and the divergence was retained in the output. Such comparisons were performed within endosymbiont groups: B. Pennsylvanicus versus B. Floridanus; Buchnera– A. Pisum versus Buchnera– S. Graminum; and Buchnera– A. Pisum versus Buchnera– Baizongia pistaciae. Divergences in endosymbionts were compared to the enterobacterial pairs E. Coli versus S. Typhimurium, and E. Coli versus P. Genomes were downloaded from NCBI in June 2004. All genomes were compared to E. Coli using RSD for ortholog detection and MultiFun-based functional assignments (detailed in Supplemental material). Rates of protein divergence along the lineages leading to B. Pennsylvanicus and B. Floridanus were compared using E. Coli as an outgroup. Relative rates were calculated as follows, with “B0” representing the common ancestor of the two Blochmannia lineages: Pairwise dN and dS values reported for certain genes [e.g., truncated genes or ORFs with apparent frameshifts within poly(A) tracts] were calculated using PAML (runmode –2) (). In order to account for the extreme base composition of endosymbiont sequences, we implemented a maximum likelihood model in which codon frequencies were calculated from the average nucleotide frequencies at the three codon positions (CodonFreq = 2). In these cases, translated Blochmannia genes were aligned to each other and to appropriate outgroups using CLUSTALX () and back-translated to nucleotide sequences using the program RevTrans (). Acknowledgments We thank Andrew McArthur, Margrethe Serres, Seth Bordenstein, Seth Kauppinen, and four anonymous reviewers for comments on the manuscript, and are grateful to Daniel Hahn, Diana Wheeler, Stefan Cover, and Diana Davidson for helpful discussion of Camponotus biology. Amy McCurley and Josh Larson contributed to the library construction, and Jeremy Brozek confirmed the B. Pennsylvanicus hisH sequence. We also thank S. Kauppinen for assistance in aligning and analyzing pseudogene sequences. Andrew McArthur, Margrethe Serres, Michael Cipriano, Sulip Goswami, Hilary Morrison, and Matt Beverly provided intellectual and technical support with genome assembly and annotation. Alessandro Romualdi provided software and support for the JENA Prokaryotic Genome Viewer, Dennis P. Wall provided access to RSD, and D. Wall and Richard Fox offered assistance implementing this program. We thank Brian Dale for permission to collect C. Pennsylvanicus for this study. This research was supported by grants to J.J.W. From NIH (R01 GM62626-01) and the NSF (DEB 0089455). Additional support was provided by the NASA Astrobiology Institute (NCC2-1054 and NNA04CC04A). Template preparation and DNA sequencing was performed in the W.M. Keck Facility for Ecological Genomics at the Marine Biological Laboratory. 1, 1, 1, 2 and 3* • 1Department of Plant Pathology, EXIM Bank Agricultural University, Chapainawabganj, Bangladesh • 2Department of Biotechnology, Bangabandhu Sheikh Mujibur Rahman Agricultural University, Gazipur, Bangladesh • 3Department of Plant Pathology, Bangabandhu Sheikh Mujibur Rahman Agricultural University, Gazipur, Bangladesh Plant growth promoting rhizobacteria (PGPR) are the rhizosphere bacteria that may be utilized to augment plant growth and suppress plant diseases. The objectives of this study were to identify and characterize PGPR indigenous to cucumber rhizosphere in Bangladesh, and to evaluate their ability to suppress Phytophthora crown rot in cucumber. A total of 66 isolates were isolated, out of which 10 (PPB1, PPB2, PPB3, PPB4, PPB5, PPB8, PPB9, PPB10, PPB11, and PPB12) were selected based on their in vitro plant growth promoting attributes and antagonism of phytopathogens. Phylogenetic analysis of 16S rRNA sequences identified these isolates as new strains of Pseudomonas stutzeri, Bacillus subtilis, Stenotrophomonas maltophilia, and Bacillus amyloliquefaciens. The selected isolates produced high levels (26.78–51.28 μg mL -1) of indole-3-acetic acid, while significant acetylene reduction activities (1.79–4.9 μmole C 2H 4 mg -1 protein h -1) were observed in eight isolates. Cucumber plants grown from seeds that were treated with these PGPR strains displayed significantly higher levels of germination, seedling vigour, growth, and N content in root and shoot tissue compared to non-treated control plants. All selected isolates were able to successfully colonize the cucumber roots. Moreover, treating cucumber seeds with these isolates significantly suppressed Phytophthora crown rot caused by Phytophthora capsici, and characteristic morphological alterations in P. Capsici hyphae that grew toward PGPR colonies were observed. Since these PGPR inoculants exhibited multiple traits beneficial to the host plants, they may be applied in the development of new, safe, and effective seed treatments as an alternative to chemical fungicides. Introduction The cucumber ( Cucumis sativus) is one of the most widely grown vegetable crops in the world, and is particularly prevalent on the Indian sub-continent. This crop is prone to massive attacks by Phytophthora capsici that causes crown rot and blight (; ). Capsici infects susceptible hosts throughout the growing season at any growth stage, and causes yield losses as high as 100% (). This pathogen has a wide host range with more than 50 plant species including Cucurbitaceae, Leguminosae, and Solanaceae (). Although fungicides can control the disease, their use is detrimental to the surrounding environment and to the viability and survival of beneficial rhizosphere microbes (;; ). Furthermore, the growing cost of pesticides and the consumer demand for pesticide-free food have led to a search for substitutes for these products. Thus, there has been a need to find effective alternatives to costly and environmentally degrading synthetic pesticides. Rhizobacteria that benefit plants by stimulating growth and suppressing disease are referred to as plant growth promoting rhizobacteria (PGPR; ). PGPR have been tested as biocontrol agents for suppression of plant diseases (), and also as inducers of disease resistance in plants (;; ). In particular, strains of Pseudomonas, Stenotrophomonas, and Bacillus have been successfully used in attempts to control plant pathogens and increase plant growth (;;;;; ). The widely recognized mechanisms of plant growth promotion by PGPR are production of phytohormones, diazotrophic fixation of nitrogen, and solubilization of phosphate. Mechanisms of biocontrol action include competition with phytopathogens for an ecological niche or substrate, as well as production of inhibitory compounds and hydrolytic enzymes that are often active against a broad spectrum of phytopathogens (;;;;;; ). Many PGPR have been shown to reduce Phytophthora crown rot occurrence on various plants. Demonstrated in vitro suppression of P. Capsici by bacterial isolates from the aerial part and rhizosphere of sweet pepper. An endophytic bacterium isolated from black pepper stem and roots, B. Megaterium IISRBP17 suppressed P. Capsici on black pepper in greenhouse assays (). Demonstrated that PGPR strains used separately or in combinations had the potential to suppress Phytophthora blight on squash in the greenhouse. Reported that some fluorescent pseudomonads isolated from different fields of East and West Azarbaijan and Ardebil provinces of Iran exhibited strong antifungal activity against P. Drechsleri and controlled crown and root rot of cucumber caused by the pathogen. However, little is known about PGPRs with the potential to suppress crown rot caused by P. Capsici on cucumber. Furthermore, the plant-growth-promoting and biocontrol efficacy of PGPR often depend upon the rhizosphere competence of the microbial inoculants (). Rhizosphere competence refers to the survival and colonization potential of PGPR (), and is thought to be highest for each PGPR strain when associated with its preferred host plant. This to some extent explains why some PGPR strains exhibiting promise as biocontrol agents in vitro have variable biocontrol efficacy in the rhizosphere of a given crop under a given set of conditions. The identification and characterization of PGPR populations indigenous to cucumber rhizospheres is therefore critical to discovery of strains that can be utilized to improve growth and Phytophthora crown rot suppression in cucumber. The objectives of the present study were to isolate bacterial strains from the cucumber rhizosphere, to characterize these isolates on the basis of morphological and physiological attributes as well as by 16S rRNA sequence analysis, and to assess the plant growth promoting effects of these isolates in vivo and their ability to suppress Phytophthora crown rot in cucumber plants. To our knowledge, this is the first report of PGPR reducing P. Capsici infection on cucumber. Materials and Methods The Study Site The experimental site was located at the Field Laboratory of the Department of Plant Pathology, Bangabandhu Sheikh Mujibur Rahman Agricultural University (BSMRAU), Gazipur, Bangladesh. The location of the site is at 24.09° N latitude and 90.25° E longitude with an elevation of 2–8 m. The study area is within the Madhupur Tract agro-ecological zone (AEZ 28). The soil used for pot experiments belongs to the Salna series and has been classified as “swallow red brown terrace soil” in the Bangladesh soil classification system, which falls under the order Inceptisol (). This soil is characterized by clay within 50 cm of the surface and is slightly acidic in nature. The pH value, cation exchange capacity (CEC) and electrical conductivity (EC) of bulk soil samples collected from the study site were 6.38, 6.78 meq 100 g -1 soil and 0.6 dS m -1, respectively. This soil contained 1.08% organic carbon (OC), 1.87% organic matter (OM), 0.10% nitrogen (N), 9 ppm phosphorus (P) and 0.20 meq 100 g -1 soil exchangeable potassium (K). Plant Material, Bacterial Isolation, and Pathogenic Organism Cucumber ( Cucumis sativus L.) variety Baromashi (Lal Teer Seed Company, Dhaka, Bangladesh) root samples were collected from the study site along with rhizosphere soil. For isolation of bacteria, 2–5 g of fresh roots were washed under running tap water and surface sterilized in 5% NaOCl for 1 min. After washing three times with sterilized distilled water (SDW), the root samples were ground with a sterilized mortar and pestle. Serial dilutions were prepared from the ground roots, and 100 μl aliquots from each dilution of 1 × 10 -6, 1 × 10 -7, and 1 × 10 -8 CFU mL -1 were spread on potato dextrose agar (PDA) plates and incubated for 2 days at 28 ± 2°C. Morphologically distinct bacterial colonies were selected for further purifications. The purified isolates were preserved temporarily in 20% glycerol solution at -20°C. The pathogen P. Capsici was provided by Prof. Yuanchao, Nanjing Agricultural University, China. Morphological and Biochemical Characterization of Bacterial Isolates Colony morphology, size, shape, color, and growth pattern were recorded after 24 h of growth on PDA plates at 28 ± 2°C as described. Cell size was observed by light microscopy. The Gram reaction was performed as described. A series of biochemical tests were conducted to characterize the isolated bacteria using the criteria of Bergey’s Manual of Systematic Bacteriology (). For the KOH solubility test, bacteria were aseptically removed from Petri plates with an inoculating wire loop, mixed with 3% KOH solution on a clean slide for 1 min and observed for formation of a thread-like mass. The motility of each isolate was tested in sulfide indole motility (SIM) medium. Using a needle, strains were introduced into test tubes containing SIM, and were incubated at room temperature until the growth was evident (). Turbidity away from the line of inoculation was a positive indicator of motility. Catalase and oxidase tests were performed as described in and, respectively. To determine whether the rhizobacterial isolates are better suited to aerobic or anaerobic environments, the citrate test was conducted according to using Simmons citrate agar medium. All experiments were done following complete randomized design (CRD) with three replications for each isolate and repeated once. Molecular Characterization of Bacterial Isolates Culture DNA was obtained using the lysozyme-SDS-phenol/chloroform method (). DNA was extracted with phenol-chloroform-isoamyl alcohol (25:24:1) and precipitated with isopropanol. The extracted DNA was treated with DNase-free RNase (Sigma Chemical Co., St. Louis, MO, USA) at a final concentration of 0.2 mg/ml at 37°C for 15 min, followed by a second phenol-chloroform-isoamyl alcohol extraction and isopropanol precipitation. Finally, the DNA pellet was re-suspended in TE buffer (10 mM Tris-HCl, 1 mM EDTA, pH 8.0), stored at -20°C, and used as template DNA in PCR to amplify the 16S rRNA for phylogenetic analysis. 16S rRNA gene amplification was performed by using the bacterial-specific primers, 27F (5 ′-AGAGTTTGATCCTGGCTCAG-3 ′) and 1492R (5 ′-GGTTACCTTGTTACGACTT-3 ′) (). PCR amplifications were performed with 1 × Ex Taq buffer (Takara Bio Inc, Japan), 0.8 mM dNTP, 0.02 units μl -1 Ex Taq polymerase, 0.4 mg ml -1 BSA, and 1.0 μM of each primer. Three independent PCR amplifications were performed at an annealing temperature of 55°C (40 s), an initial denaturation temperature of 94°C (5 min), 30 amplification cycles with denaturation at 94°C (60 s), annealing (30 s), and extension at 72°C (60 s), followed by a final extension at 72°C (10 min). The PCR product was purified using Wizard ® PCR Preps DNA Purification System (Promega, Madison, WI, USA). Purified double-stranded PCR fragments were directly sequenced with Big Dye Terminator Cycle sequencing kits (Applied Biosystems, Forster City, CA, USA) using the manufacturer’s instructions. Sequences for each region were edited using Chromas Lite 2.0. The 16S rRNA sequence of the isolate has been deposited in the GenBank database. The BLAST search program was used to search for nucleotide sequence homology for the 16S region for bacteria. Highly homologous sequences were aligned, and neighbor-joining trees were generated using ClustalX version 2.0.11 and MEGA version 6.06. Bootstrap replication (1000 replications) was used as a statistical support for the nodes in the phylogenetic trees. Bioassays for Plant Growth Promoting Traits Biological Nitrogen Fixation Nitrogenase activity of isolates was determined via the acetylene reduction assay/ethylene production assay as described in. Pure bacterial colonies were inoculated to an airtight 30 ml vial containing 10 ml nitrogen-free basal semi-solid medium, and were grown for 48 h at 28 ± 2°C. Following pellicle formation, the bottles were injected with 10% (v/v) acetylene gas and incubated at 28 ± 2°C for 24 h. Ethylene production was measured using a G-300 Gas Chromatograph (Model HP 6890, USA) fitted with a Flame Ionization Detector and a Porapak-N column. Hydrogen and oxygen were used as a carrier gas, with a flow rate of 4 kg/cm 2, and the column temperature was maintained at 165°C. The ethylene concentration calibration curve was plotted for each trial, and the viable cell numbers (cfu) of the isolate were determined. The rate of N 2 fixation was expressed as the quantity of ethylene accumulated (μmol C 2H 4 mg -1 protein h -1) based on the standard curve and peak-area percentage. Indole-3-Acetic Acid Production For detection and quantification of indole-3-acetic acid (IAA) production by bacterial isolates, isolated colonies were inoculated into Jensen’s broth (Sucrose 20 g, K 2HPO 4 1 g L -1, MgSO 4 7H 2O 0.5 g L -1, NaCl 0.5 g L -1, FeSO 4 0.1 g L -1, NaMoO 4 0.005 g L -1, CaCO 3 2 g L -1) () containing 2 mg mL -1 L-tryptophan. The culture was incubated at 28 ± 2°C with continuous shaking at 125 rpm for 48 h (). Approximately 2 mL of culture solution was centrifuged at 15000 rpm for 1 min, and a 1 mL aliquot of the supernatant was mixed with 2 mL of Salkowski’s reagent and incubated 20 min in darkness at room temperature (150 ml concentrated H 2SO 4, 250 ml distilled water, 7.5 ml 0.5 M FeCl 3.6H 2O) as described. IAA production was observed as the development of a pink-red color, and the absorbance was measured at 530 nm using a spectrophotometer. The concentration of IAA was determined using a standard curve prepared from pure IAA solutions (0, 5, 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, and 65 μg ml -1). Preparation of Bacterial Inocula for Cucumber Seed Treatment Bacterial strains were cultured in 250 ml conical flasks containing 200 ml yeast peptone broth on an orbital shaker at 120 rpm for 72 h at 28 ± 2°C. Bacterial cells were collected via centrifugation at 15000 rpm for 1 min at 4°C, and each pellet was washed twice with SDW. The bacterial pellets were suspended in 0.6 ml SDW, vortexed and used for seed treatment. Approximately 30–31 cucumber seeds were surface sterilized in 5% NaOCl for 1 min and washed three times in SDW. Dry seeds were immersed in each bacterial suspension, and the preparation was stirred frequently for 5 min. The treated seeds were spread on a petri dish and air dried overnight at room temperature. The number of bacterial cells per seed, determined via serial dilutions, was approximately 10 8 CFU/seed. Effect of Bacterial Seed Treatment on Germination and Vigour Index in Cucumber In order to determine the effect of the isolates on germination and seedling vigour, 100 seeds inoculated with each isolate were incubated in ten 9-cm petri dishes on two layers of moistened filter paper. As a control treatment, seeds treated with water instead of bacterial suspensions were also established. In order to maintain sufficient moisture for germination, 5 ml distilled water was added to the petri dishes every other day, and seeds were incubated at 28 ± 2°C in a light incubator. Germination was considered to have occurred when the radicles were half of the seed length. The germination percentage was recorded every 24 h for 7 days. Root and shoot length were measured after the seventh day. The experiment was planned as a completely randomized design with 10 replications for each isolate. V i g o u r i n d e x =% g e r min a t i o n × t o t a l p l a n t l e n g t h Effect of Bacterial Seed Treatment on Growth and Nitrogen Content in Cucumber Plants In order to test the ability of isolates to promote growth in cucumber plants, surface-sterilized cucumber seeds were inoculated with each isolate as described above. The soil from the study site described above was used as potting medium. After autoclaving twice at 24 h intervals at 121°C and 15 psi for 20 min, 180 g of the sterilized soil was placed in each sterilized plastic pot (9.5 cm × 7.0 cm size). One pre-germinated cucumber seed was sown in each pot, and plants were grown 3 weeks in a net house with watering on alternate days. After harvest, the fresh weight, dry weight, and root and shoot lengths of the plants were measured. The shoots and roots were separated and dried in an oven at 68 ± 2°C for 48 h, then ground for determination of tissue-N concentrations (). Root Colonization Root colonization by bacterial isolates was determined according to the protocol of. Roots were harvested from plants at 7, 14, and 21 days of growth. Root systems were thoroughly washed with running tap water to remove adhering soil particles, then were rinsed three times with SDW and blotted to dryness. Roots were divided into top, middle, and bottom regions, and were weighed and homogenized in SDW. Serial dilutions were prepared on PDA plates, and the number of colony-forming units (cfu) per gram root was determined after 24 h of incubation at 28 ± 2°C. Pathogenicity of P. Capsici in Cucumber For preparation of zoospore inoculum, P. Capsici was cultured on PDA plates at 18 ± 2°C for 7 days. Five-mm blocks were then cut from the culture plates and placed in petri dishes containing SDW. The petridishes were incubated in darkness at room temperature for 72 h, followed by a 1-h cold treatment at 4°C. Zoospore production was confirmed via light microscopy. In order to test the pathogenicity of P. Capsici zoospores, cucumber seedlings were planted in pots containing 0, 500, 1000, or 1500 μl zoospores/pot. As 100% mortality was found in case 1000 μl zoospores/pot, two concentrations (500 and 1000 μl) of zoospores suspension per pot were fixed. In Vitro Screening for Antagonism To test antagonism of P. Capsici by each isolate, the pathogen and bacteria were inoculated 3 cm apart on the same agar plate. Fungal growth on each plate was observed, and the zone of inhibition, if present, was determined as described in. % I n h i b i t i o n o f m y m y c e l i a l g r o w t h = X − Y X × 100 Where, X = Mycelial growth of pathogen in absence of antagonists Y = Mycelial growth of pathogen in presence of antagonists Morphologies of hyphae in the vicinity of bacterial colonies were observed under a light microscope (Meiji Techno: ML2600), and images were recorded with a digital camera (Model: Canon Digital IXUS 900 Ti). Each experiment was carried out following CRD with three replications for each isolate and repeated once. Testing the Effect of Rhizobacterial Seed Treatment on Phytophthora Crown Rot of Cucumber Cucumber seeds inoculated with each isolate were sown and grown for 7 days in sterilized plastic pots as described above. Seven-day-old seedlings were inoculated with 500 or 1000 μl zoospore suspension/pot as described in. Inoculated plants were kept inside humid chambers for 48 h. Each experiment included 12 plants per treatment with three replications. Surviving plants were counted 7 days after inoculation. Percent disease incidences (PDI) were calculated using the following formula. % p r o t e c t i o n = [ A − B A ] × 100% Where, A = PDI in non-inoculated control plants B = PDI in PGPR-treated plants. Statistical Analysis Statistical analyses were performed using SPSS (Version 17) and Microsoft Office Excel 2007. A completely randomized design was used for all experiments, with 3–12 replications for each treatment. The data presented are from representative experiments that were repeated at least twice with similar results. Treatments were compared via ANOVA using the least significant differences test (LSD) at 5% ( P ≤ 0.05) probability level. Results Strain Isolation and Biochemical and Molecular Characterization We obtained a total of 66 rhizobacterial strains from the interior of cucumber roots. Ten isolates – PPB1, PPB2, PPB3, PPB4, PPB5, PPB8, PPB9, PPB10, PPB11, and PPB12 – were selected based on their ability to produce IAA, fix N 2, and show in vitro antagonism against various pathogens in a preliminary screening. All isolates were rods producing fast-growing, round to irregular colonies with raised elevations and smooth surfaces. Reddish pigmentation was produced by PPB5, while no pigmentation was produced by other isolates (Supplementary Table S1). All 10 isolates were motile and reacted positively to the Gram staining, citrate, catalase and oxidase tests, but reacted negatively to the KOH solubility test (Table ). Phylogenetic trees constructed from 16S rRNA sequences showed that the selected isolates were mainly members of genus Bacillus, Pseudomonas, and Stenotrophomonas (Supplementary Figure ). The sequences of the isolates PPB2, PPB5, PPB8, PPB9, and PPB11 showed 99% similarity with Bacillus subtilis and were submitted to GenBank under accession numbers KJ690255, KM008605, KM008606, KM092525 and KM092527, respectively (Table ). Isolate PPB1 had 99% homology with Pseudomonas stutzeri and was submitted to GenBank under accession number KJ959616. Isolate PPB3 was identified as Stenotrophomonas maltophilia with GenBank accession number KJ959617. Isolates PPB4, PPB10, and PPB12 showed 99% sequence homology with B. Amyloliquefaciens and were submitted to GenBank under accession number KM008604, KM092526 and KM092528, respectively (Table ). Characterization for Plant Growth Promoting Traits The plant growth promoting characteristics viz., IAA production and ARA were examined with the ten selected PGPR isolates. The results of the assays are presented in Table. In the presence of tryptophan, the isolated bacteria produced IAA in concentrations between 26.78 μg mL -1 and 51.28 μg mL -1. The highest and lowest amounts of IAA were produced by strain PPB5 and PPB3, respectively (Table ). Nitrogenese activity, as determined by ARA, was not detected in PPB1 and PPB12 under the conditions tested. However, the ARA values ranged from 1.79 to 4.9 μmole C 2H 4/mg protein/h for remaining isolates. PPB2 showed the highest activity, while the lowest was recorded for PPB11 (Table ). The other isolates also reduced acetylene in significant amounts. Collectively, these results suggest that the isolates possess a number of traits associated with plant growth promotion. Germination and Vigour Index Improvement in Cucumber The effect of rhizobacterial treatment upon seed germination and vigour index of cucumber varied with different isolates. All treatments had a significant effect on the germination rate and vigour index compared to the control. The PGPR treatments increased the germination rate of cucumber seeds by 8.07–15.32% compared with the control, while the vigour index was increased by 98.62–148.05% (Figure ). In both germination rate and vigour index, the maximum increase was obtained with the PPB9 treatment. These results suggest that rhizobacterial treatment could improve the germination and vigour of cucumber seeds. Effect of plant growth promoting rhizobacteria (PGPR) treatments on seed germination, vigour and growth characteristics of cucumber seedlings grown in pots under axenic conditions. Error bars are SE from three replicates per same treatment. Data are presented as% increase in germination, vigour index, shoot length, root length, shoot fresh weight, root fresh weight, shoot dry weight, and root dry weight of PGPR-treated cucumber seedlings relative to non-treated control seedlings. The experiment was repeated twice. Plant Growth Promotion in Cucumber All isolates significantly increased the growth of cucumber compared to non-inoculated controls. Treatment with isolate PPB12 produced the maximum shoot and root lengths of 18.23 and 20.47 cm, corresponding to increases of 66.02 and 65.63% above control treatments (Figure ). However, the maximum shoot and root weight enhancement was observed in PPB8-treated plants. Treatment with isolate PPB8 produced shoot fresh and dry weights of 5.29 and 0.60 g plant -1, which were 79.32 and 100.00% higher than those of control plants. Similarly, treatment with isolate PPB8 produced root fresh and dry weights of 3.03 and 0.32 g plant -1, corresponding to increase of 91.77 and 128.57% above control treatments. N Concentration in Cucumber Plants The N content in plant roots and shoots significantly increased due to inoculation treatments with rhizobacterial isolates (Figure ). The shoot and root N content showed similar trends in response to different treatments; hence, the N content is reported as the total combined shoot and root N. The total N content in PGPR-treated plants ranged from 3.66 mg g -1 to 8.25 mg g -1 N compared with 2.57 mg g -1 N for non-inoculated control plants, a 42–221% increase in PGPR-treated plants over control plants (Figure ). The highest N content was recorded in plants grown under PPB2 followed by PPB8, PPB3, PPB9 and other treatments. Root Colonization The ability to colonize the root system is essential for rhizobacteria to be effective plant growth promoters. The root colonization assays showed that all the tested isolates successfully colonized the roots of cucumber plants as tested after only 7 days of seedling growth. The inoculated populations were even higher on 21-day-old roots. Nevertheless, the root population densities varied widely among the isolates (Figure ). The largest root populations were observed for strain PPB2, followed by PPB5 and PPB9 (Figure ). These results demonstrate specific interactions between cucumber plants and the rhizobacterial isolates. In vitro Antagonism of Phytophthora capsici All rhizobacterial isolates exhibited significant antagonistic activity against P. Capcisi on PDA. The largest inhibition zone was observed with PPB9 (90.08%) followed by PPB8 (82.05%) (Table ). Distinct morphological alterations in P. Capcisi hyphae were also detected during dual cultures with the rhizobacterial isolates. Hyphal features observed in the vicinity of bacterial colonies included irregular and excessive branching, abnormal swelling of hyphal diameters, unusually long and pointed hyphal tips, and vacuolization leading to hyphal lysis (Figure ). Suppression of Phytophthora Crown Rot in Cucumber All the selected PGPR strains showed consistent suppression of Phytophthora crown rot in the greenhouse experiments. Compared with the control, the average disease protection at 500 μl zoospore suspension ranged from 50 to 88.83% after treatment with rhizobacterial isolates, while protection at 1000 μl zoospore suspension ranged from 33.33 to 86.08% (Table ). At both inoculum rates, isolate PPB11 showed the highest disease reduction, and the lowest disease reduction was obtained with PPB5. Discussions PGPR colonizing the surface or inner part of roots play important beneficial roles that directly or indirectly influence plant growth and development (; ). In this study, 10 PGPR classified as Pseudomonas stutzeri (PPB1), B. Subtilis (PPB2, 5, 8, 9, and 11), S. Maltophilia (PPB3), and B. Amyloliquefaciens (PPB4, 10, and 12) were isolated from the rhizosphere of cucumber plants. All the isolated PGPR were gram positive and motile rods, and tested positive for the ability to utilize citrate as a carbon source. Flagellar motility and citrate utilization are both thought to play a significant role in competitive root colonization and maintenance of bacteria in roots (; ). These strains also tested positive for oxidase and catalase activity. Standard microbiology references suggest that S. Maltophilia is an oxidase-negative bacterium (). Recent data, however, suggest that some S. Maltophilia are oxidase-positive (), and this was also the case for isolate PPB3 in this study. Our catalase test results corroborate prior studies showing that B. Subtilis, Pseudomonas stutzeri, and B. Amyloliquefaciens are catalase-positive (; ). Bacillus and Pseudomonas are the most frequently reported genera of PGPR (;; ). Similarly, most isolates in this study belong to genus Bacillus. Treatment of cucumber seeds with the selected isolates significantly improved seedling emergence and growth. Several different mechanisms have been suggested for similar observations using other PGPR strains: PGPR might indirectly enhance seed germination and vigour index by reducing the incidence of seed mycoflora, which can be detrimental to plant growth (). Found that amylase activity during germination was increased in rice and legume after inoculation with PGPR. The amylase hydrolyzes the starch into metabolizable sugars, which provide the energy for growth of roots and shoots in germinating seedlings (; ). One of the most commonly reported mechanisms is the production of phytohormones such as IAA (). All the selected isolates in this study produced IAA. Similar studies have shown that IAA production is very common among PGPR (;; ). In fact, many isolates in this study produced higher IAA than previously reported strains (;; ). This is an important mechanism of plant growth promotion because IAA promotes root development and uptake of nutrients (). It has long been proposed that phytohormones act to coordinate demand and acquisition of nitrogen (). Therefore, enhanced N-content found in inoculated plants could be due to increased N-uptake by the roots caused by hormonal effects on root morphology and activity. Nitrogen fixation may also play a role in plant growth promotion. All the selected isolates in this study except PPB1 and PPB12 showed acetylene reduction activity, which is a widely accepted surrogate for nitrogenase activity and N 2-fixing potential (). However, defensible proof of N 2-fixation needs the application of 15N as tracer of soil N or as 15N 2-gas and the demonstration of significantly changed N-isotope-labeling in the plant biomass. Transfer of N between diazotrophic N-fixing rhizobacteria and the roots of several crops has been demonstrated (;;; ). It is interesting to note that in this study all isolates, including the two that demonstrated no acetylene reduction activity, enhanced the N content of cucumber. This suggests that while N 2 fixation may be an important mechanism of plant growth promotion, there may also be alternate mechanisms, like hormonal interactions and nutrient uptake or pathogen suppression, which might be more pronounced than the contribution of nitrogen fixation. Results from our study indicate that PGPR strains applied as a seed treatment significantly reduced disease severity of Phytophthora crown rot on cucumber plants. The fungal antagonists Pseudomonas stutzeri, B. Amyloliquifaciens, and S. Maltophilia were have been shown to be effective biocontrol agents in prior studies (;;;;; ). Competitive root tip colonization by PGPR strains might play an important role in the efficient control of soil-borne diseases. The crucial colonization level that must be reached has been estimated at 10 5–10 6 cfu g -1 of root in the case of Pseudomonas sp., which protect plants from Gaeumannomyces tritici or Pythium sp. Most of our selected strains were efficient colonizers of roots, since CFU counts for tested strains were more than 10 7 cfu g -1 root. However, the former study examined the root colonization by introduced bacteria under natural field soil conditions, while our study did under axenic conditions. In view of that, comparison between root colonization data obtained under these two conditions may not be accurate. Biological control of P. Capsici can also result from antibiosis by the bacteria (;;;;; ). All the selected isolates exhibited moderate to high antagonistic activity against P. Capsici in vitro, and caused clear morphological distortions such as abnormal branching, curling, swelling and lysis of the hyphae at the interaction zone in dual cultures. Excessive branching and curling accompanied by marked ultrastructural alterations including invagination of the hyphal membrane, disintegration or necrosis of hyphal cell walls, accumulation of excessive lipid bodies, enlarged and electron-dense vacuoles, and degradation of cytoplasm were potentially due to bacterial production of antibiotics and lytic enzymes (; ). These antibiotics and lytic enzymes cause membrane damage and are particularly inhibitory to zoospores of Oomycete (; ). Induced systemic resistance is most likely another mechanism by which bacteria suppress P. In the present study, we have isolated 10 new strains of PGPR from indigenous cucumber plants. These strains possessed several plant growth promoting traits as well as antifungal activity, and were found to be efficient in controlling Phytophthora crown rot of cucumber seedlings. In vitro and in vivo evidence suggest that the selected isolates benefit cucumber plants via multiple modes of action including antibiosis against phytopathogens, competitive colonization, and plant growth promotion. This reveals the potential of these strains for biofertilizer applications and commercial use as biocontrol agents in the field. However, from the estimation of a PGPR-potential to a biofertilizer application, it requires a long way of greenhouse experiments with pot filled with different type of soils and finally, field experiments to find out the optimum formulations for the inoculums. Thus, the inoculants can perform close to its optimum potential. Future studies concerning commercialization and field applications of integrated stable bio-formulations as effective biocontrol strategies are in progress. Author Contribution SI was involved in the planning and execution of the research work; collection, analysis and interpretation of the data; manuscript writing etc. Following the suggestions and directions of the Major Professor. AMA served as the Member of the Dissertation Committee of SI and was involved in the planning of the work and editing of the manuscript. AP was actively involved in the original research work, data collection, analysis as well as manuscript preparation. TI supplied the Phytophthora capsici inocula and oversaw the sequence work of the bacterial isolates and related description in the manuscript. MMH served as the Major Professor of SI and was involved in the research design and planning; analysis and interpretation of data; drafting as well as critical revision of the work for intellectual content. All authors approve the final version of the manuscript for publication and agrees to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. Conflict of Interest Statement The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. Acknowledgments The authors would like to acknowledge the financial assistance from University Grant Commissions through Research Management Committee of Bangabandhu Sheikh Mujibur Rahman Agricultural University, Bangladesh. The authors also wish to thank Prof. Islam for his help obtain 16S rRNA sequences of the tested bacterial isolates. Supplementary Material The Supplementary Material for this article can be found online at: FIGURE S1| Phylogenetic tree of 16S rRNA gene sequences showing the relationships among the isolates isolated from cucumber rhizosphere. The data of type strains of related species were from GenBank database. • References. Keywords: PGPR, plant growth promotion, IAA production, biological nitrogen fixation, antagonism, Phytophthora capsici, disease suppression Citation: Islam S, Akanda AM, Prova A, Islam MT and Hossain MM (2016) Isolation and Identification of Plant Growth Promoting Rhizobacteria from Cucumber Rhizosphere and Their Effect on Plant Growth Promotion and Disease Suppression. Doi: 10.3389/fmicb.2015.01360 Received: 16 August 2015; Accepted: 16 November 2015; Published: 02 February 2016. Reviewed by:, University of Texas at Arlington, USA, Sun Yat-sen University, China Copyright © 2016 Islam, Akanda, Prova, Islam and Hossain. This is an open-access article distributed under the terms of the. The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms. *Correspondence: Md. 
Xtreme Download Manager ( xdman) is a powerful download manager for Linux, which is developed in Java programing language. It can increase download speeds up to 500% and is an alternative for the windows IDM ( Internet Download Manager). It can be integrated with any internet browser such as Firefox, Chrome, Opera and many more and supports pause and resume functionalities while downloading files. Apr 4, 2015 - 5 min - Uploaded by ZigZag TecHow to Fix when you have Java SE 6 Runtime Issue on Mac OSx Yosemite. Click the link below. Update Release Notes summarize changes made in all 1.6.0 update releases. Note that bug fixes are cumulative, that is, bug fixes in previous update versions are. Oracle’s Java runtime software is required to run Java applets on websites and desktop software written in the Java programming language. When installing Java. Don’t Miss: XDMAN has the following features: • It is highly portable therefore runs on any O.S with Java SE 6, no need for installation. • It downloads files at maximum possible speed. • It has a sophisticated dynamic file segmentation algorithm,data compression & connection reuse. • It can download FLV, MP4, HTML5 videos from YouTube, MySpaceTV, Google Video or videos from many other sites. • It can capture download from any browser (Firefox, Chrome, Internet Explorer, Opera, Safari or any other program trying to download a file from Internet). • It supports HTTP, HTTPS, FTP protocols with authentication, proxy servers, cookies, redirection etc. • It can aslo resumes broken/dead downloads caused by connection problem, power failure or session expiration. • It has a built in YouTube downloader, HTTP Traffic monitor and batch downloader. • It can also be configured to perform automatic antivirus checking, system shutdown on download completion. To use XDMAN, you need to have Java installed on your Linux system. You can check if Java is installed or not by typing java -version in command line. $ java -version java version '1.8.0_45' Java(TM) SE Runtime Environment (build 1.8.0_45-b14) Java HotSpot(TM) 64-Bit Server VM (build 25.45-b02, mixed mode) If incase, Java not installed, you can install it using your default system package manager yum or apt. Install Xtreme Download Manager in Linux To install most recent stable version of Xtreme Download Manager ( XDM) in Linux distributions such as Ubuntu, Debian, Linux Mint, Fedora, etc. First you need to download a zipped file using wget utility and install it by running the following commands. On 32-bit Systems $ wget $ tar -xvf xdm-jre-32bit.tar.xz $ cd xdm $ ls -l $./xdm On 64-bit Systems $ wget $ tar -xvf xdm-jre-64bit.tar.xz $ cd xdm $ ls -l $./xdm Note: Alternatively, you can double click on xdm file to launch it and create application shortcut from XDM Menu -> File -> Create application shortcut on Desktop as shown below. How about you show us an example of where it increased the speed of a download by 500%? Or how about you explain what techniques it uses to increase the download speed. There are a couple of possibilities: 1) download different parts of the same file from multiple sources (ie, torrents) maybe works with CDN websites 2) download different parts of the same file from the same source, maybe works when the source limits download speed per connection rather than per source IP address. Not really seeing how this is going to help many people, and not prepared to install it. Also, a note that XDM is the X windows Desktop Manager, so you should be careful to call it XDMan instead of XDM (unless you are purposefully trying to be confusing to generate more interest). When you do software review, you not only give what you have experienced while using it but also include what the developers say about how it works. The speed boost of 500% may seem to be over exaggerated but that is how the developers think the algorithm they used works. Users work in different environments and network speeds or data connections speeds vary therefore, showing how that actually works can not be possible sometimes especially the way you want it to be. The best idea would be to let the developers know about your concerns and show you how it actually works, boosting speeds up to 500%. We do not want to give information just to win support but to enlighten users, share knowledge, bring out our experiences and what has been stated. This is the XDM project homepage:, you can find more form there or even contact the developers. Advertisement Oracle’s Java runtime software is required to run on websites and desktop software written in the Java programming language. When you install Java, there are a few things you should consider, especially regarding security. Java is used by an ever-decreasing number of websites and is a frequent target of attacks. Most people could remove Java and not notice a difference. If you do use Java, you should be aware of the security problems and take proper precautions. You’ll also need to know whether you need the Java Runtime Environment (JRE) or the Java Development Kit (JDK). You May Not Need To Install Java Do you use a specific website or program that requires Java? If not, you don’t actually need it installed. Java just allows you to run software written in Java, and you may be surprised by how few websites and programs actually require Java. If you’re not sure whether you need Java, try going without it for a while. You may not notice the difference. As we’ll detail later, there are good reasons not to have Java installed — if you can help it. Even (formerly OpenOffice.org), doesn’t require Java for most of its functionality. JDK The offers the Java Runtime Environment, also known as the JRE. This is the one you probably need. It includes the basic software that lets you run Java applets and desktop applications on your computer. There’s also the Java Development Kit, also known as the JDK. This is what you need if you want to. Some development-related software, including the, also requires the JDK on your system. If you need the JDK, you’ll have to. The JDK also includes the JRE, so you only have to install one of them. Security Problems The elephant in the room when it comes to installing Java is security. Browser plugins – particularly Oracle’s Java and Adobe’s Flash and – are major targets. Java is a prime target because it’s installed on so many computers and exploits will work across multiple browsers and operating systems. Keeping Java updated doesn’t fully alleviate this problem – just having Java installed increases your browser’s attack surface. Update Java Often If you need Java installed, you’ll want to update it often. By default, Java checks for updates once every month – not a very reassuring default setting for a program that’s so frequently exploited. You can fix this, though. To do so, open the Control Panel from the Start menu, click the Programs category, and click the Java icon. Some Software Requires Older Versions Depending on the the software you use, you may not be able to run the latest, secure versions of Java. Some websites and applications specify a specific version of Java and force you to use an outdated, vulnerable version of Java. This is why it’s possible to have multiple versions of Java installed on the same system, although Oracle recommends against this. Oracle maintains if you need to, while noting that they’re full of security holes and vulnerable to attack. If you must run an older version of Java, make sure you have an installed, ask the application vendor or website for an update, and remove the old version of Java as soon as possible. Installing Java Applets Can Be Dangerous Web browsers and plugins such as Flash isolate web content from your computer. A website with a Flash-based video player can’t break out of your browser and tamper with your computer (barring security vulnerabilities). Java does the same thing for most applets, which it runs by default – but it also allows applets to prompt you for full permissions. Office 2016 Permanent Activator Ultimate v1.2 adalah activator permanen yang pertama ada untuk mengaktifkan microsoft office 2016 anda secara permanen. Anda yang sudah menggunakan office 2016 namun masih belum mengaktifkan program tersebut secara permanen, maka sudah seharusnya anda mencoba menggunakan Office 2016 Permanent Activator yang admin bagikan kali ini. Dengan menggunakan activator ini maka office 2016 milik anda akan menjadi aktif secara permanen. Cara kerja activator yang satu ini adalah dengan mengupdate secara otomatis MAK Key Office 2016 yang terinstal di pc atau laptop anda. Untuk dapat menggunakan Office 2016 Permanent Activator ini, anda memerlukan koneksi internet di pc atau laptop anda. Jadi bagi anda yang ingin menggunakan activator ini untuk komputer yang selalu offline, sepertinya harus menunda dulu. Karena ini membutuhkan koneksi internet untuk mengupdate MAK Key tersebut. Kemudian pc atau laptop anda juga harus terinstal Net Framework 4.5 ke atas. To get started: • Click the NI Downloader link above. • Your browser will download the NI Downloader launcher for your software (~150kb). Tabtight professional, free when you need it, VPN service. 
• Once NI Downloader launcher has been downloaded, launch the executable. • The NI Downloader will automatically initiate the download of your software's standalone installer. • When the download of the standalone installer is complete, launch it and follow the onscreen prompts to complete the installation of your software. Note: Features of the NI Downloader include the ability to pause/resume download, or automatically resume download if unintentionally interrupted. $266 millions a year in dividends, taxed at only 15% (because our beloved job creators are worth it). That's $226 millions a year after taxes. Every second of every day (and night), your fearless leader earns as much as a a fast food worker gets for an hour of scrubbing toilets and cleaning burnt grease. No doubt there are pinko commies here who will scream 'social justice' but let's face it, business acumen like that of Steve doesn't come cheap. It takes a special kind of individual to come up with ideas like the aQuantive acquisition or the Kin. 
• maggio 31, 2017 a 9:00 pm|. I usually do not leave a response, but after looking at a few of the comments here Embed Code| Intelicom. I do have a couple of questions for you if you do not mind. Is it only me or does it give the impression like a few of these responses appear like they are written by brain dead folks? And, if you are posting on additional social sites, I would like to follow you. Could you make a list of the complete urls of all your social pages like your linkedin profile, Facebook page or twitter feed? • giugno 3, 2017 a 8:48 am|. 女性の毛って見られてます。つるすべ肌ってもはや当然。エステ脱毛の選び方をレポートしました。家での一時しのぎはありえないって話です。素肌を傷つけるし、黒ずみの原因ですよ。でもって、トラブルと美肌に戻すのに期間も必要です。それにね、傷んだ肌を目にするのは落ち込みます。めげますよね。だからプロの出番。サロンの脱毛でトラブルもなく美しいスベスベ肌を目指せます。太い毛でも大丈夫ですよ。太い毛でも料金加算なんてありえないのが嬉しいですよね。何より、毛深いほどにきれいになる体感が早いのが嬉しいところです。毛深いのを見られるのがハズカシイって方がいますが、スタッフの方は気にしてないですよ。なぜって何十人ものユーザーを相手にしてるから考えてる時間なんてあるわけないです。気遣いは必要ないってことです。肌がすべすべだとバッチリ胸をはって水着が着られるのが嬉しいです。脱毛が完了すると絶対にいいです。割チケで申込むと激安に脱毛できるからぜひトライしてみてくださいね。勧誘が不愉快だという人もいますが、最近ではそんなにキツくないです。そんなことは気にしないでOKでしょう。毛深いのを家で処理してる手間がせつないですよー。簡単にきれいになりましょう。 • giugno 9, 2017 a 4:56 pm|. サラリーマンで副業って、なんだか難しい感覚ですよね?実際に副業を始めようと思ってみても、具体的なノウハウやビジネスモデルのやり方が不明だと手をつけることは出来るわけありませんよね。 初心者におすすめの副業は、転売ビジネスです。転売ビジネスとは、ビジネスの基礎であるモノの売買でマージンを稼ぐビジネスモデルです。なんでこれが良いのかというと、取り組みやすいからです。 転売ビジネスにおいて、例えばホームセンターでセール品などを購入し、ヤフオクなどのネットショップで買った時の金額を超える金額でモノを売り払えばよいのです。購入金額と売却金額の差分と送料などを減らした金額が、純利益になるのです。 そして、転売ビジネスはアマゾンなどのネットショップに登録するだけでマイ店舗をお手軽に開業できるので、ゴリ押ししているのです。リアルにお店を開こうものなら初期費用でかなり持っていかれますので。やらない手はありません。 他にもサイトアフィリエイトやクラウドワーキングなど、難易度が難しいけど大きく稼げるものや誰でもできるけど金額に見合っていないなど、多種多様なビジネスがあります。あなたの生活スタイルにあった副業を見つけて、後悔のない人生を送りましょう。 その他副業についてはこちらにたくさん記載しています。知りたい人はぜひ見てみてください。 • giugno 10, 2017 a 12:56 pm|. Good day I am so delighted I found your weblog, I really found you by accident, while I was browsing on Google for something else, Nonetheless I am here now and would just like to say thank you for a marvelous post and a all round exciting blog (I also love the theme/design), I don’t have time to go through it all at the minute but I have book-marked it and also added your RSS feeds, so when I have time I will be back to read a lot more, Please do keep up the great work. • luglio 3, 2017 a 9:26 pm|. Gallery: hard reset samsung galaxy t679 t-mobile move balance case t-mobile move cena cherry mobile t11 applications t11 cherry mobile features cherry mobile t11 free download them. I think it's really useful and important for a writer to experience these sorts of things — and your descriptions really bring out the magic in what other people might see as mundane, ordinary. Y marianne costa pdf tirada de cartas egipcias gratis como leer el tarot cruz celta leer las cartas del tarot en linea tarot gratis de. From Wikipedia: “Walter Matthau was initially offered the role of the judge, but demanded a fee of US$1 million. The producers balked at meeting his price and signed Alan Arkin instead for a modest $150,000. Arkin was replaced by Morgan Freeman when the studio decided to change the judge’s ethnicity from Jewish to African-American in order to moderate criticism of the film’s racial politics, and dialogue was added to have the judge give the final denouncement towards the manipulative actions of the main characters. Murray Abraham, who has a significant part in the film, chose to not be credited over a contract dispute.”. Jcibal: Laker fans are soft. I have been posting about this. Other teams have seas of identical tee-shirts, + we have a crowd “waiting to be entertained”. Posting on a blog doesn’t hurt the team. Going to the game and being silent does. When I go to games – I scream and yell + try to get others to do the same. Muddywood: I understand your post, however plugging the hole in the 2nd team just opens one up on the first team. MB should not have to make that trade off, but you are correct that he does. Too bad the average American can't read this and lacks the cognizance to be pissed off, if they had an ounce of self respect and moral vision, they would not be so easily manipulated by Politicians. At least Teddy had some testicular fortitude, America's demise started in 1913 with the creation of the Federal Reserve, that is when the Government was usurped, true evil and it's type of spineless men have been pulling the levers ever since. Thanks again Daniel for saying it like it is, at least someone knows how to speak the truth, you are the antidote to stupidity and arrogance in Washington. You were dyslexic.' The truth is, mentoring helps me and the kids that I mentor:) La~~incognita, you are on a roll today. You have some funny stuff with you there girl.:)Thanks for the kind words StillaPanther2. We are trying to hold it down here on the East Coast. Black Diaspora I have been wondering about that too. Where are all the white Obama supporters when all of this kind of crap goes down? Let's hope that they have been equally outraged at the bull s*#t, and that they have been sending e-mails and calls to the corporations involved. SomeChick FUCK YOU! One of those idiots that don’t speak ENGLISH!!!!!!!!!!!!!!!!!!! I would never have had any issues with Sprint if the FUCKWADS who answer the customer service line spoke English or at the least understood it!! I KNOW what I signed up for and SPRINT failed to tell me that they changed the plans this year! SO FUCK YOU BITCH! Your such a chicken crap you don’t even put your real name!But you can cuss people out who are in the right eh? Vas faire foutre a la cache!!!!!!!!!!!!!!!!!! Your comments are well taken, BUT enough is enough! It is time for the Republican Party to get some backbone and take up the fight. Trying to be civil, just, and right against people who will do and say anything is a losing battle. Have we not learned anything from the last 2 elections!! We need candidates who are not afraid to get in a liar’s face! I plan to leave the Republican Party and register as an Independent. The RP has gotten so weak that they are equally responsible for the demise in this “Great” country, which is coming. It is a sad, sad day in American history. Gunslinger–I think the only way to fight this is by 1. Asserting our rights as Americans and 2. Most politicians from the city councilman to the president need money from big lobbyists. But what they need even more are voters.A politician doesn't start out being a politician. He/she starts out as a public servant and from there a politician addicted to power. We can take the power away by simply voting the bad ones out of office and showing that we mean what we say by lobbying.Politicians need to become public servants again or given the royal boot. Eu nu am acordat o atentie prea mare apei de gura niciodata,recunosc insa aceasta postare ma pune pe ganduri. Multumim pentru sfaturi!Dar la capitolul paste de dinti, o recomandare, pls? Sensodyne, de pilda, isi merita renumele? Sa fie un produs mai bun decat Colgate, de pilda? Am cumparat Sensodyne (Fluoride) care este usor mentolata, curata bine dar nu-mi dau seama sa aiba ceva in plus fata de Colgate sau Blend-a-Med (alte doua branduri care imi plac). Poate la cap. Compozitie, Sensodyne sa difere in sens pozitiv? Nu ma pricep. MIchael,This isn’t working still. One issue is that I cannot find this in the php to change. The price will display once you choose a variant but I would like it to display the price before you choose your variant. Also, I can’t get it to display the new price on the product listing page.3.6 Replace the following snippet “$sale_price = get_product_meta($product_id, ‘price’, true);” on line #1811 in /wp-content/plugins/wp-e-commerce/wpsc-includes/product-template.php$regular_price = wpsc_calculateprice( $product_id, null, false ). File Viewer Lite supports: • Windows 10 • Windows 8 • Windows 7 • Windows Vista • Windows XP File Viewer Lite is a universal file opener that allows you to open any file on your Windows PC. Simply drag and drop any file or folder onto the application icon or program window. File Viewer Lite can open PDFs, word processing documents, spreadsheets, and many other common file types in their native view. It also can play dozens of audio and video file formats. If you have a unknown file, File Viewer Lite can open it! In the rare case that File Viewer Lite does not natively support the file format, you can still view the contents of any file in text or hexadecimal view. ✓ Follow File Viewer on or on. ✓ Learn how to use File Viewer Lite by reading the. ✓ File Viewer Lite for Windows contains file type information from. ✓ This software uses code licensed under the and its source can be downloaded. Other platforms: You can also download and. Do you have documents on your computer that you can't open? Windows File Viewer Lite is the solution! It is a free application that allows you to open and view over 150 different file types. File Viewer Lite supports all major multimedia formats and many less common audio and video formats as well. It includes playback controls so you can play movies and sound files. You can view uncommon file types in File Viewer Lite, then copy the images or text to the Windows clipboard. You can then paste the data into another application. File Viewer Lite displays detailed information about every file you open. It even shows hidden metadata, like EXIF data saved with digital images and PDF author information.  
Generally, a download manager enables downloading of large files or multiples files in one session. Many web browsers, such as Internet Explorer 9, include a download manager. Stand-alone download managers also are available, including the Microsoft Download Manager. If you do not have a download manager installed, and still want to download the file(s) you've chosen, please note: • You may not be able to download multiple files at the same time. In this case, you will have to download the files individually. (You would have the opportunity to download individual files on the 'Thank you for downloading' page after completing your download.) • Files larger than 1 GB may take much longer to download and might not download correctly. • You might not be able to pause the active downloads or resume downloads that have failed. The Microsoft Download Manager solves these potential problems. It gives you the ability to download multiple files at one time and download large files quickly and reliably. It also allows you to suspend active downloads and resume downloads that have failed. Microsoft Download Manager is free and available for download now. • NOTE: The Microsoft PowerPoint Viewer is being retired in April, 2018. At that time, the Viewer will no longer be available for download and will no longer receive security updates. Microsoft recommends using these applications to view PowerPoint files: • On Windows 10 PCs, download the free PowerPoint Mobile application from the • On iOS devices, download the free PowerPoint for iOS application from the Apple Store • On Android devices, download the free PowerPoint for Android application from the Google Play store • Try Office 365 for free for 1 month, which gives you access to the full versions of Word, Excel, PowerPoint, Outlook and more and can be installed on up to 5 PCs or Macs, plus up to 5 tablets and 5 phones. • On Windows 7 or Windows 8/8.1 PCs, upload the file to OneDrive and view it for free using PowerPoint Online PowerPoint Viewer lets you view full-featured presentations created in PowerPoint 97 and later versions with full fidelity. This viewer also supports opening password-protected Microsoft PowerPoint presentations. Jan 14, 2008 Open, view, and print Excel workbooks, even if you don't have Excel installed. This download is a replacement for Excel Viewer 97 and all previous Excel. You can view and print presentations, but you cannot edit them in the PowerPoint Viewer. The following section on Font Components amends the license terms for the PowerPoint Viewer and must accompany any permitted redistribution of the PowerPoint Viewer: Font Components You may use the fonts that accompany the PowerPoint Viewer only to display and print content from a device running a Microsoft Windows operating system. Additionally, you may do the following: • Embed fonts in content as permitted by the embedding restrictions in the fonts • When printing content, temporarily download the fonts to a printer or other output device You may not copy, install or use the fonts on other devices. Note: If no end user license agreement is presented during installation or the first run of PowerPoint Viewer, by downloading this software you agree that the software is subject to the terms of the end user license agreement that you already accepted with your previous installation of PowerPoint Viewer, with Microsoft as the licensor. The following Microsoft PowerPoint features are not supported by this viewer: • Information Rights Management (IRM) presentations. • Running macros, programs, or opening linked or OLE objects. Supported Operating System Windows 7, Windows Server 2003 R2 (32-Bit x86), Windows Server 2003 R2 x64 editions, Windows Server 2008, Windows Vista Service Pack 1, Windows Vista Service Pack 2, Windows XP Service Pack 3 • • Memory: • 256 MB RAM or more • 270 MB of available hard disk space • Display: 1024x768 or higher resolution monitor • Additional: Graphics hardware acceleration requires a DirectX 9.0c graphics card with 64 MB or more video memory. Product functionality and graphics may vary based on your system configuration. • This download works with presentations created in the following Microsoft Office programs: • Microsoft PowerPoint 2010 • Microsoft Office PowerPoint 2007 • Microsoft Office PowerPoint 2003 • Microsoft PowerPoint 2002 • Microsoft PowerPoint 2000 • Microsoft PowerPoint 97. • To install this download: • Download the file by clicking the Download button (above) and saving the file to your hard disk. • Double-click the PowerPointViewer.exe program file on your hard disk to start the setup program. • Follow the instructions on the screen to complete the installation. Instructions for use: • Once you have installed this download, from the Start menu under All Programs, open Microsoft PowerPoint Viewer. Note PowerPoint Viewer registers with the.ppt,.pptx,.pptm,.pot,.potx,.potm,.pps,.ppsx and.ppsm file extensions only if a version of PowerPoint is not installed on your computer. If registered, double-clicking on these file types will launch PowerPoint Viewer. To remove this download: • On the Start menu, point to Settings, and then click Control Panel. • Double-click Add/Remove Programs. • In the list of currently installed programs, select Microsoft PowerPoint Viewer, and then click Remove or Add/Remove. • Click Yes or OK to confirm that you want to remove the program. To remove the download file • Delete the file PowerPointViewer.exe. Update (2016)– As Microsoft has closed down Nokia firmware repository (on December 31, 2016), softwares like Navifirm, Nokia Care Suite, Nokia Data Package Manager are not working anymore. Navifirm developer has promised to fix this issue in the new version of Navifirm to be released soon. Until then you can use to download Nokia phones software files. For latest updates on Nokia/Symbian you may refer to Using Navifirm Plus to download Nokia firmware files Back in the golden days of Nokia it was pretty easy to get Nokia phone’s firmware files but things are different now. Nokia has been decimated by Microsoft and everything related to Nokia is behaving weirdly these days including their software download servers. Nokia had already closed down the doors of their software servers for general public and only authorized Nokia personnel now have server access via login. Due to these restrictions, many firmware-downloading softwares stopped working. Fortunately some are still working thanks to the changes made by their developers, Navifirm Plus is one such software. Before this server shutdown episode Navifirm was completely free to use but now it’s a “ freemium” software – you have to pay (or “donate” as the developer calls it) a nominal amount to get a personal account to unlock all the features. However one can still use Navifirm without paying money with some restrictions of course. So you have to enter the product code of your phone to download its software files. Product codes of Nokia phones refer to a 7 character long code consisting of numerals or a combination of numbers and alphabets for example product code of Nokia N8 (Indian variant) is 0599255 Product codes mostly start with a zero. Phone’s product code is usually printed at the sticker found under the battery and/or at the retail box. It is also displayed by some phones when *#0000# is dialed. Note down your phone’s product code and enter it in the “ Search in Nokia Care Suite” box (at the top-left corner of Navifirm) and press enter at the keyboard Navifirm will search Nokia Care servers for available software files After a while Navifirm Plus will load the product code info (in products, releases, variants panes) and the software files (in the last pane). Navifirm Plus Options Look at the software files in the last pane – some files are pre-selected by default (which are necessary) while some are not (which are optional). Usually you do not have to mark or un-mark any of these as all the essential files are marked and the unmarked files are optional. However you can click “ Mark all” button to select all files just to be on safer side. The memory card content file (usually the last and the biggest file in the list), if present, can be safely left out because it is completely optional and takes too much time to download (due to its size). Now un-check the “ Use ProductCode as final path” box and then click “ Browse” button to set the location where Navifirm will save phone software files – better create a new folder at the “Desktop” and name it as RM-xxx where last three xxx refer to your phone type for example RM-596 for Nokia N8 (you can find out yours at the sticker below phone battery or by dialing *#0000#). Sep 16, 2010. The Nokia N8 is a fantastic mobile movie maker, and once you've mastered our tips for creating Hollywood-grade HD home movies, you'll want to refine the. Oct 20, 2009. Nokia Map Loader can download less data over-the-air. You can add map data and navigator voices to Ovi Maps on your mobile before you travel. Download and. 
This way it will be easy for you to copy and paste this folder at a specific location (required for flashing the phone). It is completely optional though and you can save software files anywhere you want including the default location pre-set in NaviFirm. Now click “ Download” button – Navifirm Plus will start downloading the files – wait for the process to compete (downloading time will depend upon the phone model and your internet connection speed). Note – Sometimes Navifirm gives error at start up or while loading/downloading software files. These errors are due to Nokia server issues and can be solved by closing and re-launching Navifirm a few times. If still unsolved – try launching Navifirm after some time (say after 1 hour) as server overload or other issues are resolved. I hope this post clarifies everything related to the use of Navifirm Plus. If you still have any question, feel free to drop in a comment. Besides you may refer to which describes some Navifirm alternatives which can be used to download Nokia firmware files. • Nokia 8 is the first flagship Android phone of the new Nokia • Coolpad Cool Play 6 is the cheapest 6 GB RAM phone in India at Rs.14,999 • WhatsApp stops working in Symbian S60 phones • Asus ZenFone AR, the first 8 GB RAM phone goes on sale • UMDIGI Z Pro is the first Android phone to have “Live Photo” function • OnePlus 5 caught for manipulating Benchmark scores by XDA devs • Galaxy Note 8 is the first dual-camera Samsung flagship • A request to our readers – Symbian related links in old blog-posts may not work now. Please report such broken links via comments or Contact Us page so that we can update the same. Thanks in advance. 
Mahabharat karan vs arjun tamil movies free download, mahabharat karan vs arjun hd video songs download, mahabharat karan vs arjun download youtube video, mahabharat karan vs arjun video songs download, mahabharat karan vs arjun tamil songs free download, hindi video song download, mahabharat karan vs arjun hindi video songs free download, tamil play, mahabharat karan vs arjun bollywood movies 2015, mahabharat karan vs arjun mp4 video, mahabharat karan vs arjun mp4 song, mahabharat karan vs arjun tamil hd video songs, mahabharat karan vs arjun new tamil movies free download. Jatt Jaguar song from Mubarakan, A song with the Punjabi taste from a Bollywood Movie, starring Arjun Kapoor, Anil Kapoor, Ileana D’Cruz and Athiya Shetty. Tags: Karan Arjun Hd1080p Song Video Songs, Karan Arjun Hd1080p Song bollywood movie video, Karan Arjun Hd1080p Song video Download, mp4 Karan Arjun Hd1080p Song hindi movie songs download, Karan Arjun Hd1080p Song all video download, Karan Arjun Hd1080p Song Hd Video Songs, Karan Arjun. Mobile toplist for mobile web sites. We have over 2000 registered sites.  |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
